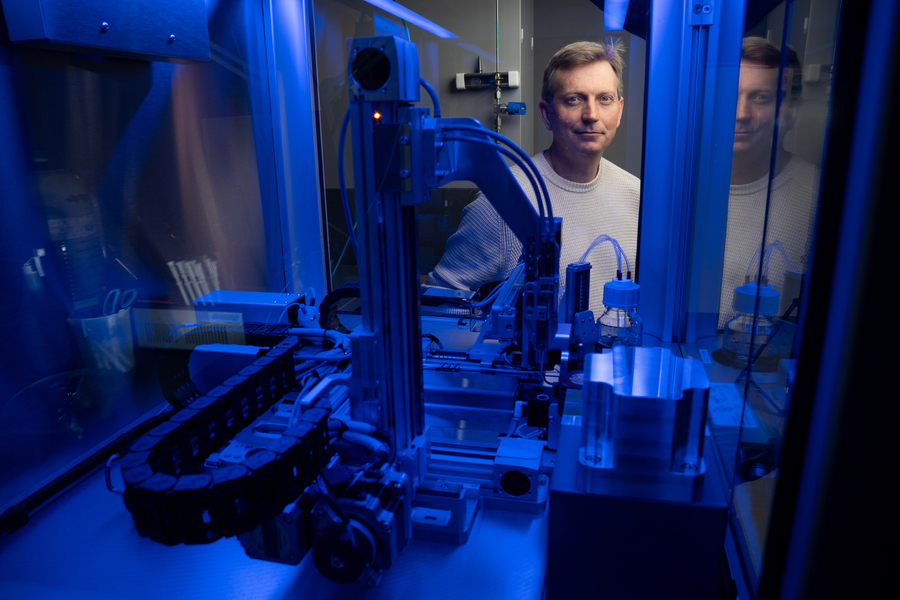
His studies have shed light on the assembly instructions that govern ribosomes, the critical protein-building machines of the cell.
Anne Trafton | MIT News
May 5, 2026
Ribosomes, the cellular machines that assemble proteins, are made from dozens of proteins and RNA molecules. Putting all of those pieces together is a complex puzzle — one that MIT Associate Professor Joey Davis PhD ’10 revels in trying to solve.
Understanding how these structures form and later break down could help researchers learn more about how disruptions of these fundamental processes can lead to disease. But, as Davis points out, it’s also an interesting biological question.
“Our long-term goal is to really understand how the natural world assembles these huge complexes rapidly and efficiently. It’s a fundamentally interesting question to think about how these things get put together,” he says.
His work has helped reveal that unlike building a house, which happens in a prescribed sequence of steps — pouring the foundation, building the frame, putting on the roof, then doing electrical and plumbing work — ribosomes can be assembled in a more flexible way. Cells can even skip an assembly step and then come back to it later.
“In these natural systems, it seems like the assembly pathways are much more dynamic and flexible,” he says. “It appears that evolution has selected pathways that aren’t strictly ordered in the way we would think about an assembly line, where you always put in one component, then the next, and then the next. We’re excited to understand the selective advantages of such approaches.”
A love of discovery
Davis’ interest in how things are put together developed early in life, inspired by his father, a carpenter who framed houses. During the mid-1980s, the family moved from Colorado to Southern California, where his father worked in construction during a housing boom there.
“I was always interested in building things, which I think probably came from being around my dad and other builders,” Davis says.
As an undergraduate at the University of California at Berkeley, where he majored in computer science and biological engineering, Davis’ interests turned toward smaller scales, in the realm of cells and molecules. During his junior year, he started working in the lab of chemistry professor Michael Marletta, who studies molecular-level biological interactions.
In the lab, Davis investigated how enzymes that contain heme are able to preferentially bind to either oxygen or nitric oxide, two gases that are very similar in structure. That work kindled a love of studying the natural world and pursuing discoveries in fundamental science.
“Being in the Marletta lab and seeing students and postdocs that were really passionate about these problems had a big impact on me,” Davis says. “The goal was to understand the fundamentals of how molecular discrimination works, and the idea of discovery for the sake of discovery was thrilling.”
After graduating from Berkeley, Davis spent another year working in Marletta’s lab, and then a year working odd jobs, before heading to MIT to pursue a PhD in biology. There, he worked with Professor Bob Sauer, now emeritus, who studied the relationship between protein structure and function, with a particular focus on the molecular machines that degrade or remodel proteins.
Davis’ thesis research centered on enzymes called AAA proteases, which remove damaged proteins from cellular membranes and send them to cell organelles that break them down. In addition to studying the structure and function of the proteases, Davis worked on ways to engineer them to tag specific proteins for destruction.
That work led him into synthetic biology, which he used to develop genetic parts that drive production of proteins of interest. Some of those parts ended up being used by the biotech startup Ginkgo Bioworks, where Davis took a job as a senior scientist after graduating.
Working at Ginkgo Bioworks allowed Davis to stay in Boston while his partner finished her PhD. The couple then moved back to California, where Davis worked as a postdoc at Scripps Research, which was home to one of the first direct electron detection cameras for cryo-electron microscopy (cryo-EM). These detectors allow researchers to generate structures with near atomic resolution. At Scripps, Davis began using them to study ribosomes as they were being assembled.
Peering into the ribosome
After joining the MIT faculty in 2017, Davis continued his work on ribosomes and assembled a lab group that includes students from a variety of backgrounds who work together to develop new ways to explore biological phenomena.
“I have a mix of method developers and biologists in the group, and the work from each of them informs each other,” Davis says. “My lab goes back and forth between building sets of tools to answer biological questions, and then as we’re answering those questions, it motivates the next generation of tool development.”
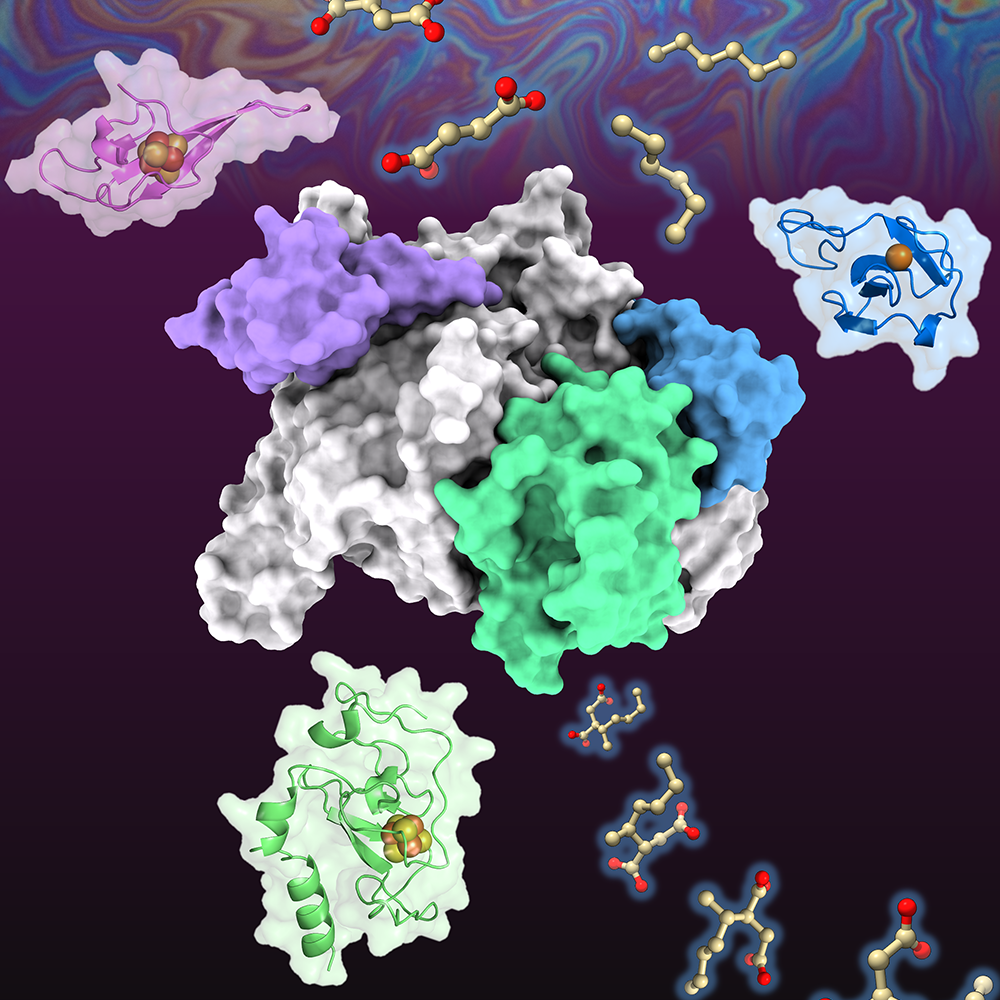
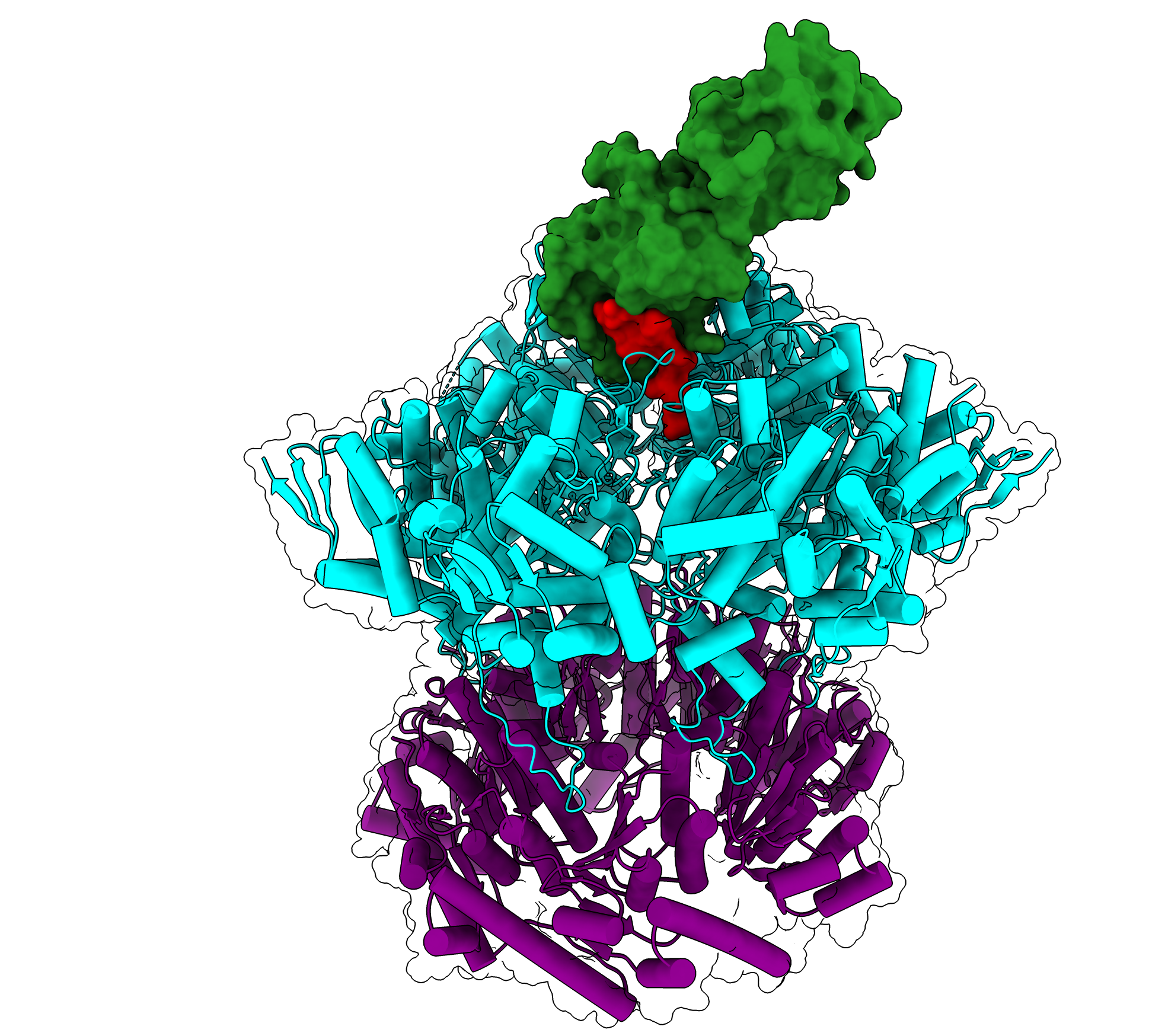
During ribosome assembly, RNA molecules fold themselves into the correct shapes, creating docking sites for proteins to attach. Then, more RNA molecules come in and fold themselves into the structure.
“It’s a beautifully coupled process by which the cell folds hundreds of RNA helices and binds on the order of 50 proteins, and it does it in two minutes from start to finish. E. coli does this 100,000 times per hour, and it’s amazing how rapid and efficient the process is,” Davis says.
Cryo-EM allows scientists to capture this process in minute detail. It can be used to take hundreds of thousands of two-dimensional images of ribosome samples frozen in a thin layer of ice, from different angles. Computer algorithms then piece together these images into a three-dimensional representation of the ribosome.
To gain insight into how ribosomes are assembled, researchers can stall the process at different points and then analyze the resulting structures. In 2021, Davis’s lab developed a new method called CryoDRGN, which uses neural networks to analyze cryo-EM data and generate the full ensemble of structures that were present in the sample.
This work has shown that when certain steps of ribosome assembly are blocked, many different structures result, suggesting that the assembly can occur in a variety of ways.
In future work, Davis aims to dramatically increase the throughput of cryo-EM to generate datasets of protein structures that could help improve the AI-based models that are now used to predict protein structures.
“There are still huge swaths of sequence space that these models are very poor at predicting, but if we could collect data on those sequences en masse, that could potentially serve as key training data for a next-generation protein structure prediction method that could fill out that space,” he says.