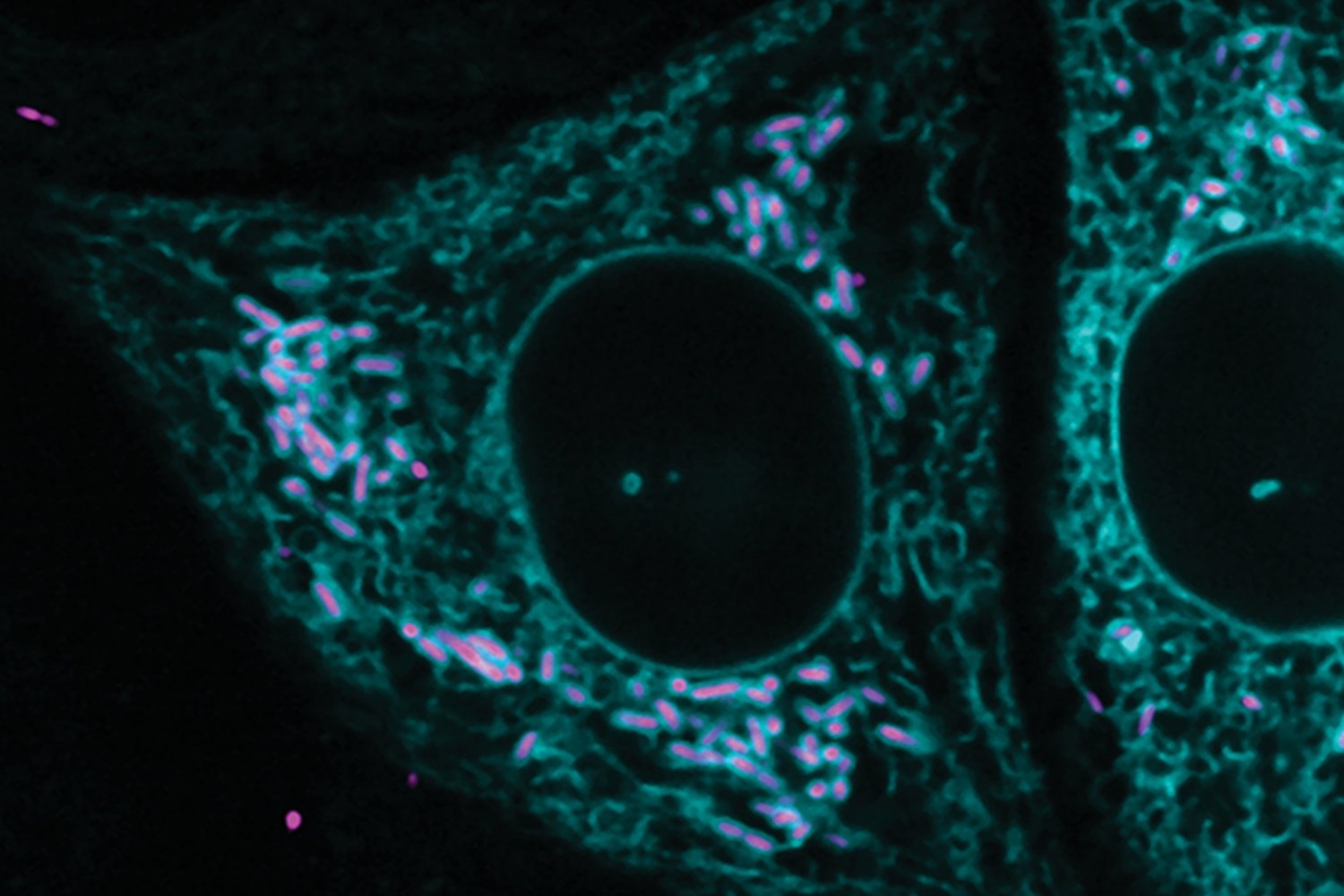
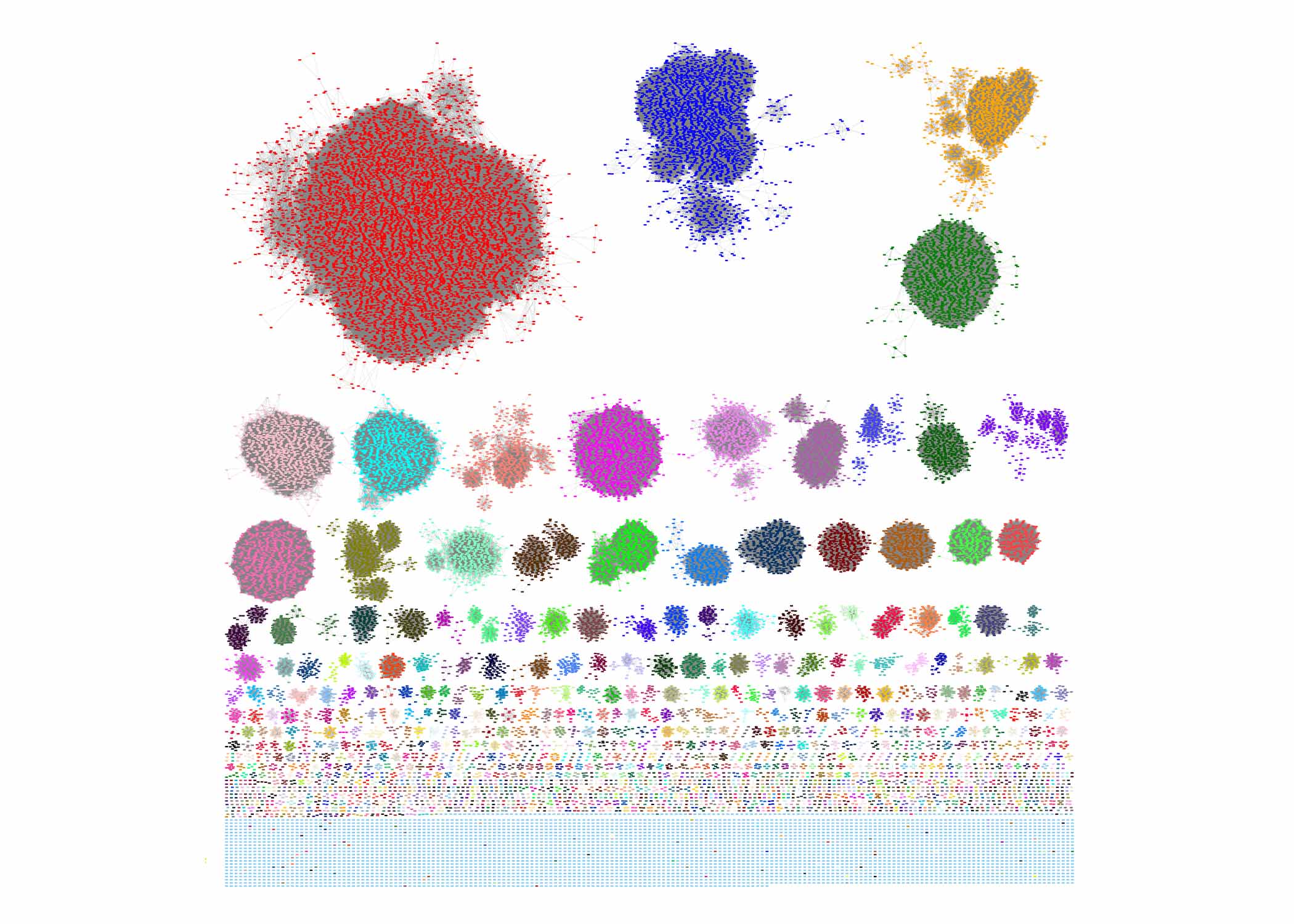
One of the ways that cells ensure proteins end up where they're needed is creating them at that location, through a process called localized translation. New research from the Weissman Lab has expanded our understanding localized translation at mitochondria and sheds light on the organizational principles of genes and the proteins they encode.
Greta Friar | Whitehead Institute
August 27, 2025
Now, Weissman, who is also a professor of biology at the Massachusetts Institute of Technology and an HHMI Investigator, and postdoc in his lab Jingchuan Luo have expanded our knowledge of localized translation at mitochondria, structures that generate energy for the cell. In a paper published in Cell on August 27, they share a new tool, LOCL-TL, for studying localized translation in close detail, and describe the discoveries it enabled about two classes of proteins that are locally translated at mitochondria.
The importance of localized translation at mitochondria relates to their unusual origin. Mitochondria were once bacteria that lived within our ancestors’ cells. Over time the bacteria lost their autonomy and became part of the larger cells, which included migrating most of their genes into the larger cell’s genome in the nucleus. Cells evolved processes to ensure that proteins needed by mitochondria that are encoded in genes in the larger cell’s genome get transported to the mitochondria. Mitochondria retain a few genes in their own genome, so production of proteins from the mitochondrial genome and that of the larger cell’s genome must be coordinated to avoid mismatched production of mitochondrial parts. Localized translation may help cells to manage the interplay between mitochondrial and nuclear protein production—among other purposes.
How to detect local protein production
For a protein to be made, genetic code stored in DNA is read into RNA, and then the RNA is read or translated by a ribosome, a cellular machine that builds a protein according to the RNA code. Weissman’s lab previously developed a method to study localized translation by tagging ribosomes near a structure of interest, and then capturing the tagged ribosomes in action and observing the proteins they are making. This approach, called proximity-specific ribosome profiling, allows researchers to see what proteins are being made where in the cell. The challenge that Luo faced was how to tweak this method to capture only ribosomes at work near mitochondria.
Ribosomes work quickly, so a ribosome that gets tagged while making a protein at the mitochondria can move on to making other proteins elsewhere in the cell in a matter of minutes. The only way researchers can guarantee that the ribosomes they capture are still working on proteins made near the mitochondria is if the experiment happens very quickly.
Weissman and colleagues had previously solved this time sensitivity problem in yeast cells with a ribosome-tagging tool called BirA that is activated by the presence of the molecule biotin. BirA is fused to the cellular structure of interest, and tags ribosomes it can touch—but only once activated. Researchers keep the cell depleted of biotin until they are ready to capture the ribosomes, to limit the time when tagging occurs. However, this approach does not work with mitochondria in mammalian cells because they need biotin to function normally, so it cannot be depleted.
Luo and Weissman adapted the existing tool to respond to blue light instead of biotin. The new tool, LOV-BirA, is fused to the mitochondria’s outer membrane. Cells are kept in the dark until the researchers are ready. Then they expose the cells to blue light, activating LOV-BirA to tag ribosomes. They give it a few minutes and then quickly extract the ribosomes. This approach proved very accurate at capturing only ribosomes working at mitochondria.
The researchers then used a method originally developed by the Weissman lab to extract the sections of RNA inside of the ribosomes. This allows them to see exactly how far along in the process of making a protein the ribosome is when captured, which can reveal whether the entire protein is made at the mitochondria, or whether it is partly produced elsewhere and only gets completed at the mitochondria.
“One advantage of our tool is the granularity it provides,” Luo says. “Being able to see what section of the protein is locally translated helps us understand more about how localized translation is regulated, which can then allow us to understand its dysregulation in disease and to control localized translation in future studies.”
Two protein groups are made at mitochondria
Using these approaches, the researchers found that about twenty percent of the genes needed in mitochondria that are located in the main cellular genome are locally translated at mitochondria. These proteins can be divided into two distinct groups with different evolutionary histories and mechanisms for localized translation.
One group consists of relatively long proteins, each containing more than 400 amino acids or protein building blocks. These proteins tend to be of bacterial origin—present in the ancestor of mitochondria—and they are locally translated in both mammalian and yeast cells, suggesting that their localized translation has been maintained through a long evolutionary history.
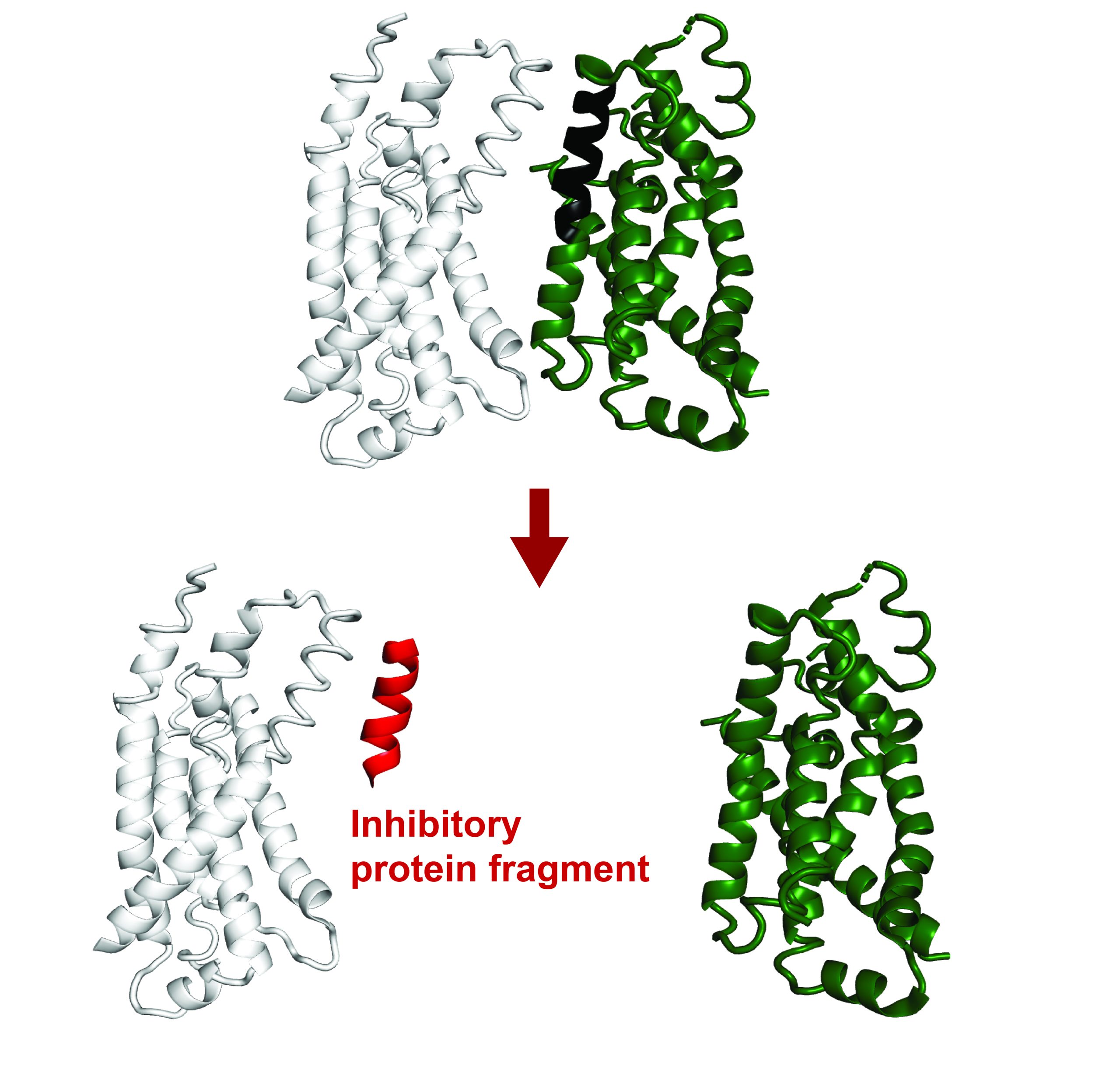
Like many mitochondrial proteins encoded in the nucleus, these proteins contain a mitochondrial targeting sequence (MTS), a zip code that tells the cell where to bring them. The researchers discovered that most proteins containing an MTS also contain a nearby inhibitory sequence that prevents transportation until they are done being made. This group of locally translated proteins lacks the inhibitory sequence, so they are brought to the mitochondria during their production.
Production of these longer proteins begins anywhere in the cell, and then after approximately the first 250 amino acids are made, they get transported to the mitochondria. While the rest of the protein gets made, it is simultaneously fed into a channel that brings it inside the mitochondria. This ties up the channel for a long time, limiting import of other proteins, so cells can only afford to do this simultaneous production and import for select proteins. The researchers hypothesize that these bacterial-origin proteins are given priority as an ancient mechanism to ensure that they are accurately produced and placed within mitochondria.
The second locally translated group consists of short proteins, each less than 200 amino acids long. These proteins are more recently evolved, and correspondingly, the researchers found that the mechanism for their localized translation is not shared by yeast. Their mitochondrial recruitment happens at the RNA level. Two sequences within regulatory sections of each RNA molecule that do not encode the final protein instead code for the cell’s machinery to recruit the RNAs to the mitochondria.
The researchers searched for molecules that might be involved in this recruitment, and identified the RNA binding protein AKAP1, which exists at mitochondria. When they eliminated AKAP1, the short proteins were translated indiscriminately around the cell. This provided an opportunity to learn more about the effects of localized translation, by seeing what happens in its absence. When the short proteins were not locally translated, this led to the loss of various mitochondrial proteins, including those involved in oxidative phosphorylation, our cells’ main energy generation pathway.
In future research, Weissman and Luo will delve deeper into how localized translation affects mitochondrial function and dysfunction in disease. The researchers also intend to use LOCL-TL to study localized translation in other cellular processes, including in relation to embryonic development, neural plasticity, and disease.
“This approach should be broadly applicable to different cellular structures and cell types, providing many opportunities to understand how localized translation contributes to biological processes,” Weissman says. “We’re particularly interested in what we can learn about the roles it may play in diseases including neurodegeneration, cardiovascular diseases, and cancers.”
Luo et al. “Proximity-specific ribosome profiling reveals the logic of localized mitochondrial translation.” Cell, August 27, 2025. https://doi.org/10.1016/j.cell.2025.08.002