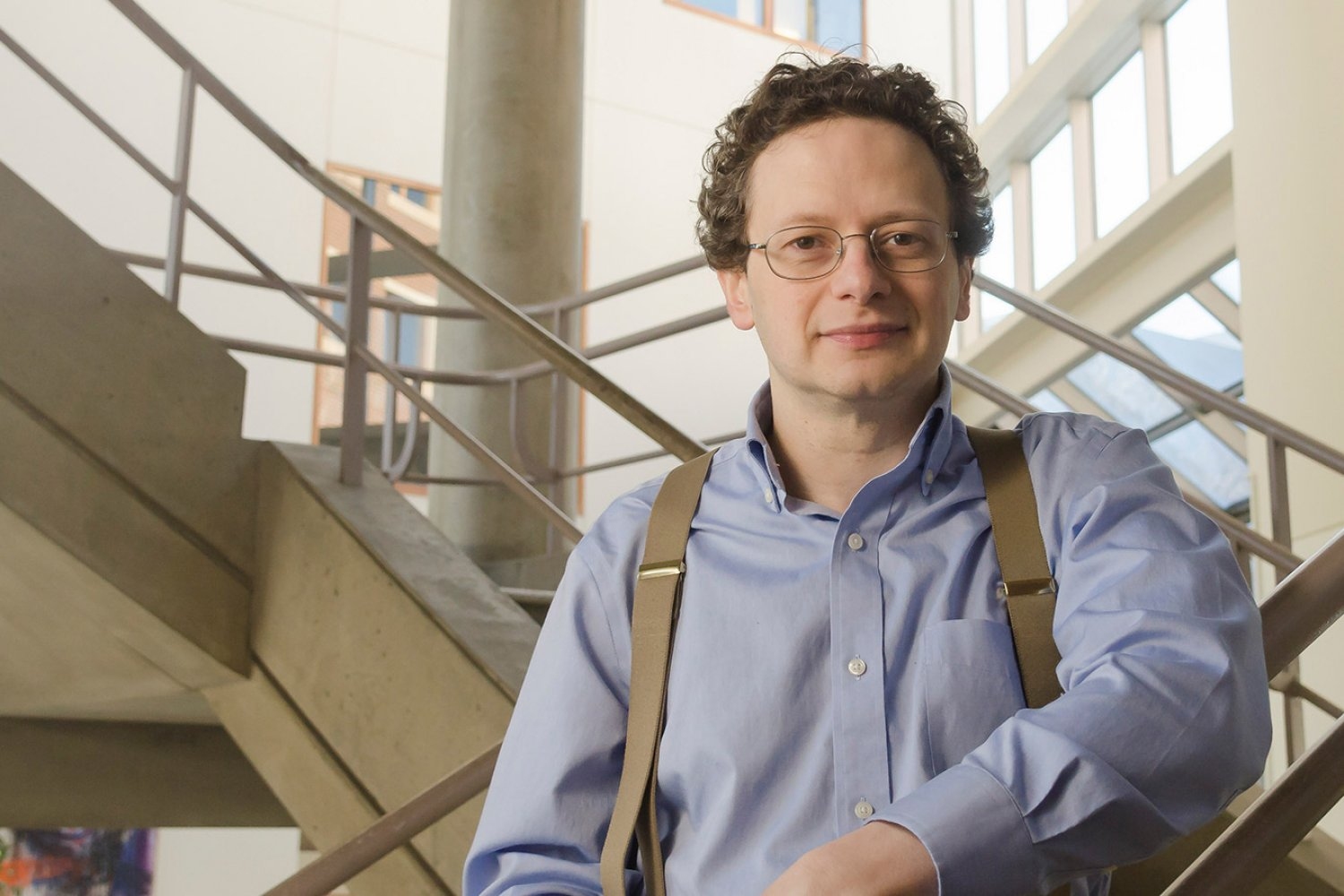
Stuart Levine ’97, director of MIT’s BioMicro Center, keeps departmental researchers at the forefront of systems biology.
Samantha Edelen | Department of Biology
March 19, 2025
As director of the MIT BioMicro Center (BMC), Stuart Levine ’97 wholeheartedly embraces the variety of challenges he tackles each day. One of over 50 core facilities providing shared resources across the Institute, the BMC supplies integrated high-throughput genomics, single-cell and spatial transcriptomic analysis, bioinformatics support, and data management to researchers across MIT.
“Every day is a different day,” Levine says, “there are always new problems, new challenges, and the technology is continuing to move at an incredible pace.” After more than 15 years in the role, Levine is grateful that the breadth of his work allows him to seek solutions for so many scientific problems.
By combining bioinformatics expertise with biotech relationships and a focus on maximizing the impact of the center’s work, Levine brings the broad range of skills required to match the diversity of questions asked by researchers in MIT’s Department of Biology.
Expansive expertise
Biology first appealed to Levine as an MIT undergraduate taking class 7.012 (Introduction to Biology), thanks to the charisma of instructors Professor Eric Lander and Amgen Professor Emerita Nancy Hopkins. After earning his PhD in biochemistry from Harvard University and Massachusetts General Hospital, Levine returned to MIT for postdoctoral work with Professor Richard Young, core member at the Whitehead Institute for Biomedical Research.
In the Young Lab, Levine found his calling as an informaticist and ultimately decided to stay at MIT. Here, his work has a wide-ranging impact: the BMC serves over 100 labs annually, from the the Computer Science and Artificial Intelligence Laboratory and the departments of Brain and Cognitive Sciences; Earth, Atmospheric and Planetary Sciences; Chemical Engineering; Mechanical Engineering; and, of course, Biology.
“It’s a fun way to think about science,” Levine says, noting that he applies his knowledge and streamlines workflows across these many disciplines by “truly and deeply understanding the instrumentation complexities.”
This depth of understanding and experience allows Levine to lead what longtime colleague Professor Laurie Boyer describes as “a state-of-the-art core that has served so many faculty and provides key training opportunities for all.” He and his team work with cutting-edge, finely tuned scientific instruments that generate vast amounts of bioinformatics data, then use powerful computational tools to store, organize, and visualize the data collected, contributing to research on topics ranging from host-parasite interactions to proposed tools for NASA’s planetary protection policy.
Staying ahead of the curve
With a scientist directing the core, the BMC aims to enable researchers to “take the best advantage of systems biology methods,” says Levine. These methods use advanced research technologies to do things like prepare large sets of DNA and RNA for sequencing, read DNA and RNA sequences from single cells, and localize gene expression to specific tissues.
Levine presents a lightweight, clear rectangle about the width of a cell phone and the length of a VHS cassette.
“This is a flow cell that can do 20 human genomes to clinical significance in two days — 8 billion reads,” he says. “There are newer instruments with several times that capacity available as well.”
The vast majority of research labs do not need that kind of power, but the Institute, and its researchers as a whole, certainly do. Levine emphasizes that “the ROI [return on investment] for supporting shared resources is extremely high because whatever support we receive impacts not just one lab, but all of the labs we support. Keeping MIT’s shared resources at the bleeding edge of science is critical to our ability to make a difference in the world.”
To stay at the edge of research technology, Levine maintains company relationships, while his scientific understanding allows him to educate researchers on what is possible in the space of modern systems biology. Altogether, these attributes enable Levine to help his researcher clients “push the limits of what is achievable.”
The man behind the machines
Each core facility operates like a small business, offering specialized services to a diverse client base across academic and industry research, according to Amy Keating, Jay A. Stein (1968) Professor of Biology and head of the Department of Biology. She explains that “the PhD-level education and scientific and technological expertise of MIT’s core directors are critical to the success of life science research at MIT and beyond.”
While Levine clearly has the education and expertise, the success of the BMC “business” is also in part due to his tenacity and focus on results for the core’s users.
He was recognized by the Institute with the MIT Infinite Mile Award in 2015 and the MIT Excellence Award in 2017, for which one nominator wrote, “What makes Stuart’s leadership of the BMC truly invaluable to the MIT community is his unwavering dedication to producing high-quality data and his steadfast persistence in tackling any type of troubleshooting needed for a project. These attributes, fostered by Stuart, permeate the entire culture of the BMC.”
“He puts researchers and their research first, whether providing education, technical services, general tech support, or networking to collaborators outside of MIT,” says Noelani Kamelamela, lab manager of the BMC. “It’s all in service to users and their projects.”
Tucked into the far back corner of the BMC lab space, Levine’s office is a fitting symbol of his humility. While his guidance and knowledge sit at the center of what elevates the BMC beyond technical support, he himself sits away from the spotlight, resolutely supporting others to advance science.
“Stuart has always been the person, often behind the scenes, that pushes great science, ideas, and people forward,” Boyer says. “His knowledge and advice have truly allowed us to be at the leading edge in our work.”