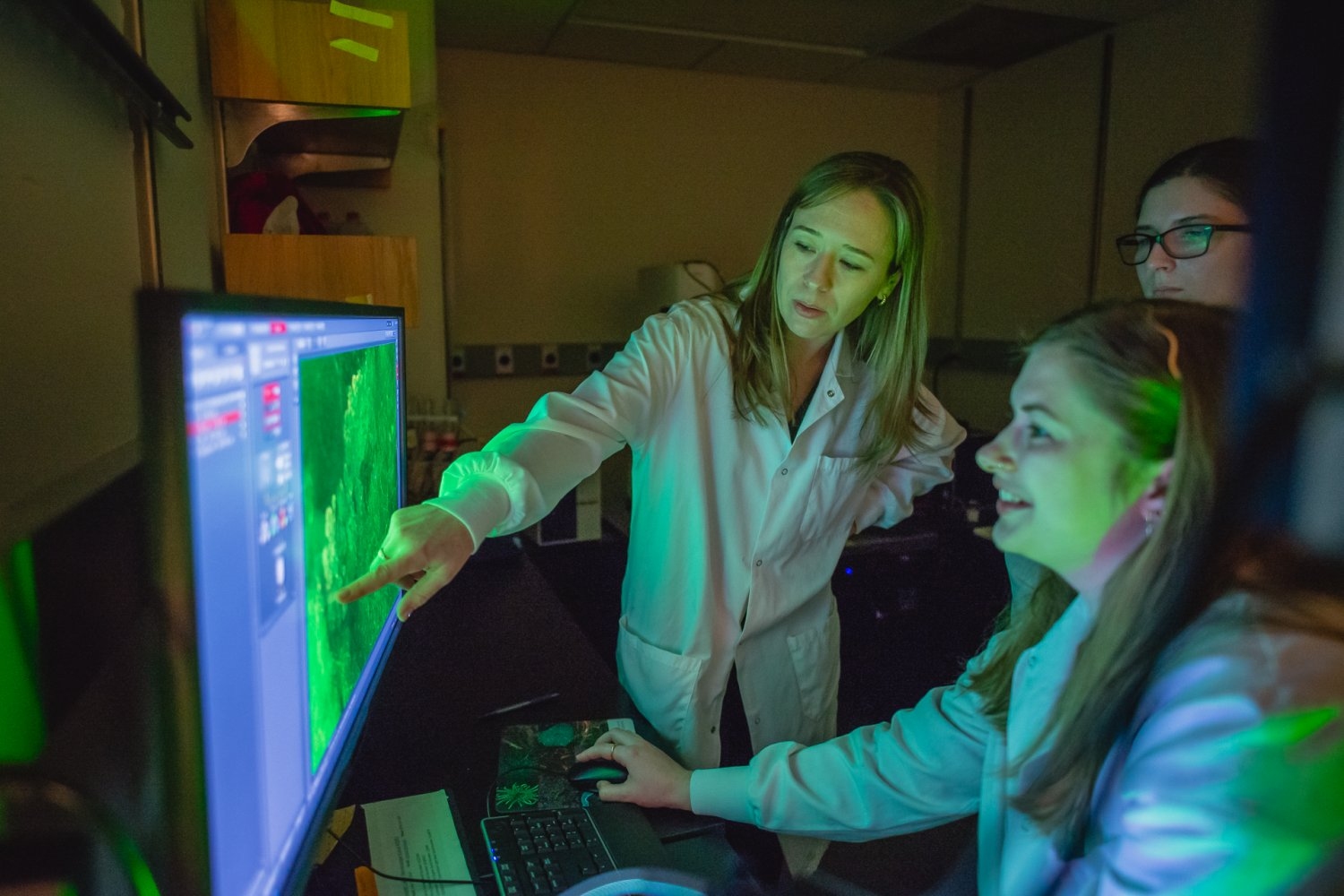
“We cannot be effective scientists if we are unhappy or unhealthy outside of the lab,” says “Committed to Caring” honoree Sara Prescott.
Leila Hudson | Office of Graduate Education
March 27, 2026
Professor Sara Prescott embodies the kind of mentorship every graduate student hopes to find: grounded in scientific rigor, guided by kindness, and defined by a deep commitment to well-being. Her approach reflects a simple but powerful belief that transformative mentorship is not only about advancing research, but about cultivating confidence, belonging, and resilience in the next generation of scholars.
A member of the 2025–27 Committed to Caring cohort, Prescott exemplifies the program’s spirit, which honors faculty who go above and beyond in nurturing both the intellectual and personal development of MIT’s graduate students.
Prescott is the Pfizer Inc. – Gerald D. Laubach Career Development Professor in the MIT departments of Biology and Brain and Cognitive Sciences, and an investigator at the Picower Institute for Learning and Memory. Her research addresses fundamental questions in body-brain communication, with a focus on lung biology, early-life adversity, women’s health, and the impacts of climate change on respiratory health.
A culture of compassion
Prescott’s mentoring philosophy begins with a focus on professional sustainability. “We cannot be effective scientists if we are unhappy or unhealthy outside of the lab,” she says.
She pushes back against what she sees as an unhelpful narrative in academia. “There’s this idea that you must choose between a successful PhD or having a personal life. This is a false dichotomy, and a problematic attitude.” Instead, she reminds her mentees that “graduate school is a marathon, not a sprint,” encouraging them to place importance not only on their research, but also on their mental and physical well-being.
This set of values shines through within her lab climate as a whole. Students describe support for flexible scheduling and mental health leave, a willingness to reimburse meals during late-night lab sessions, and encouragement during stretches of experimental failure. In addition to these more technical supports, nominators also shared stories of Prescott engaging in the smaller details: prioritizing connection for her students, celebrating their milestones, organizing lab retreats, and fostering a culture where people feel valued beyond their productivity.
Students recognize Prescott as a safe haven within the often complex and challenging world of research. Joining Prescott’s lab was a turning point for one student who was recovering from a damaging prior mentorship experience. They arrived uncertain, struggling to trust faculty and questioning whether they belonged in science at all. Prescott met them with empathy and professionalism, offering patience and trust not just in their work, but in them as a person. They describe steady support that, over time, helped them “fall back in love with science” and envision a future they had nearly abandoned.
Prescott draws inspiration from the mentorship she received early in her career. As a trainee, she had mentors who helped her believe that she could succeed. Now in a mentoring role herself, she does her best to pass this sense of confidence on to her advisees.
She is intentional about creating space where students can grow without fear. From their very first meetings, one nominator wrote, Prescott emphasized that “graduate school is a place for learning and curiosity.” They never felt judged for not knowing something; instead, they were encouraged to ask questions, share ideas, and take intellectual risks. That environment, the student explained, allowed them to grow into their scientific identity with confidence.
Prescott reinforces this message often. Success, she tells students, grows from effort, learning, and persistence, rather than from fixed traits. When working with students, she does her best to reframe failure as part of the process, emphasizing its importance within the scientific journey. Through these avenues, she cultivates a lab culture where nominators are challenged to think boldly while feeling genuinely supported, and where her students are seen not only as researchers, but as whole people.
Advocacy beyond the bench
Prescott’s commitment to caring extends well beyond day-to-day lab work. Her nominators relate that she actively supports her students’ professional development, encouraging them to pursue writing projects, certificates, internships, leadership roles, and community engagement.
Nominators also highlight Prescott’s focus on supporting underserved communities within the field as a whole. Students highlight her involvement with Graduate Women in Biology (GwiBio), where she volunteered as a speaker for the “Glass Shards” series. Her talk “Failure as the Path to Success,” in which she candidly shared pivots and setbacks in her own career, was described as one of the organization’s most impactful sessions.
Her dedication to inclusion is equally evident in her mentorship of scholars whose role in her lab is more temporary. She welcomes international visiting scholars, temporary lab techs, and undergraduate interns in the MIT Summer Research Program. When one intern encountered barriers at their home institution, Prescott ensured they had a continued research home in her lab at MIT. These additional resources allowed them to complete their undergraduate thesis and graduate on time from their university.
Prescott says that she views mentorship as an evolving practice, regularly soliciting feedback from her students. Effective leadership, in her view, grows from mutual trust and open communication.
For many nominators, Prescott’s impact extends beyond their careers. “She has taught me what positive and supportive mentoring relationships look like,” one student reflected. “When I think about the type of mentor I want to be, I hope I can emulate the ways in which she supports and guides nominators to develop their scientific independence and confidence.”
In lifting up the people behind the science as thoughtfully as the science itself, Sara Prescott demonstrates that the most enduring legacy of a mentor is not only the discoveries from their lab, but the composure and courage their advisees carry forward.