A summer intensive using microscopy to study a unique type of yeast was a dream come true for BSG-MSRP-Bio student Adryanne Gonzalez.
Lillian Eden | Department of Biology
September 11, 2025
For Adryanne Gonzalez, studying yeast using microscopy at MIT this summer has been a dream come true.
“Whatever world we’re living in, there’s an even smaller one,” Gonzalez says. “Knowing and understanding the smaller one can help us learn about the bigger stuff, and I think that’s so fascinating.”
Gonzalez was part of the Bernard S. and Sophie G. Gould MIT Summer Research Program in Biology, working in the Lew Lab this summer. The program offers talented undergraduates from institutions with limited research opportunities at their home institutions the chance to spend 10 weeks at MIT, where they gain experience, hone skills, and create the types of connections with potential collaborators and future colleagues that are critical for success in academia.
Gonzalez was so excited about the opportunity that she didn’t apply for any other summer programs.
“I really wanted to work on becoming more independent in the lab, and this program was research-intensive, and you get to lead your own project,” she says. “It was this or nothing.”

The fun of science & the rigors of mentoring
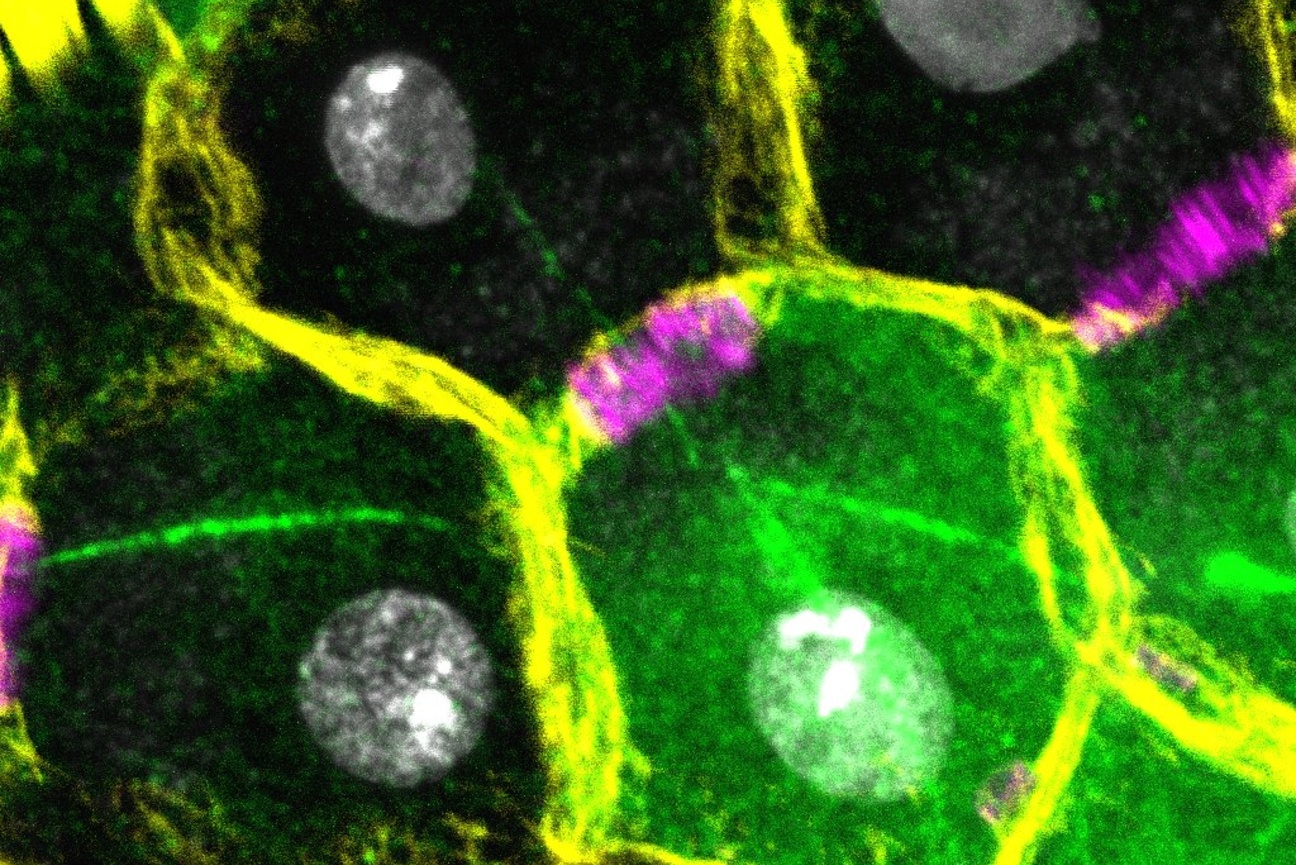
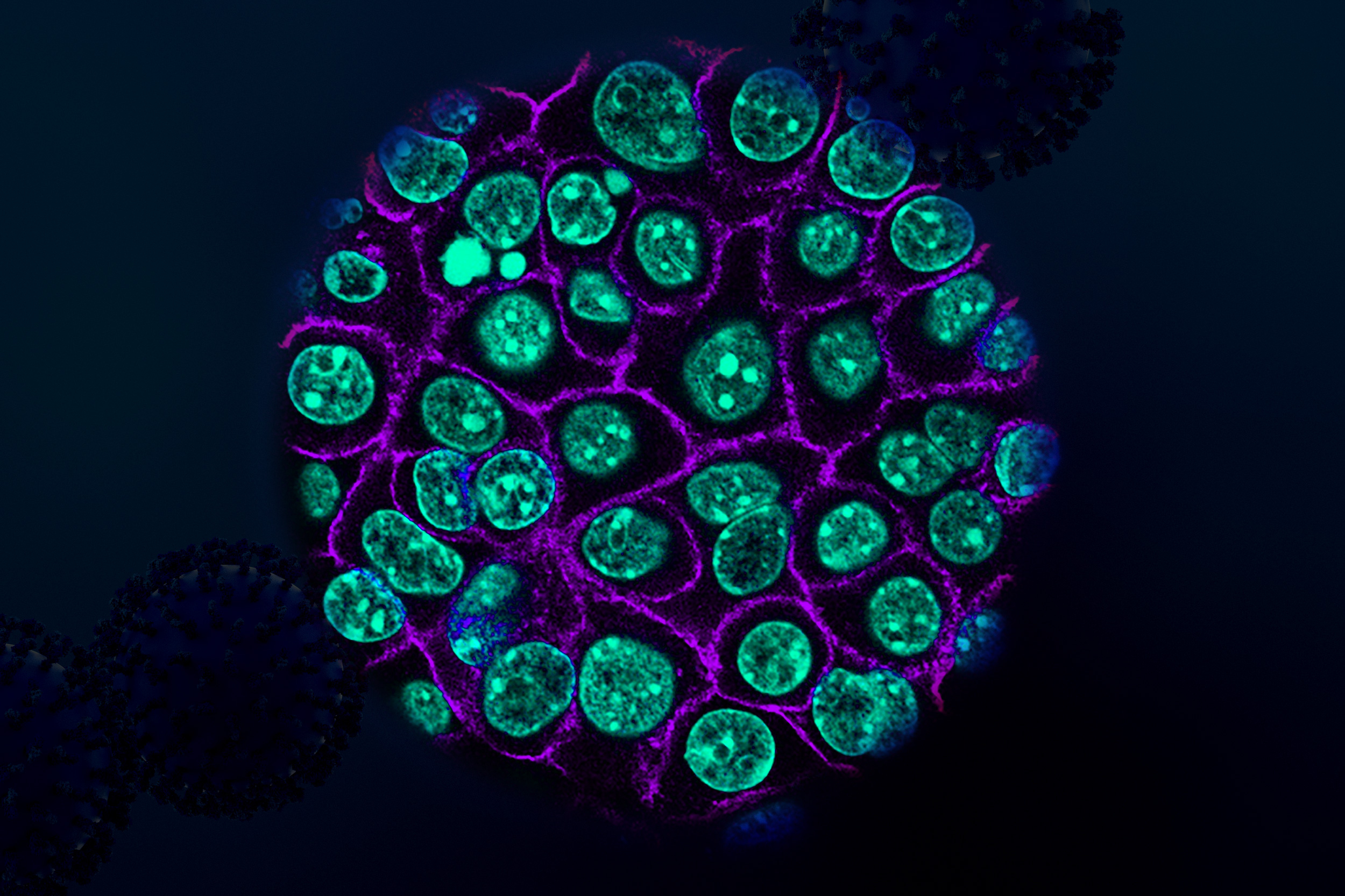
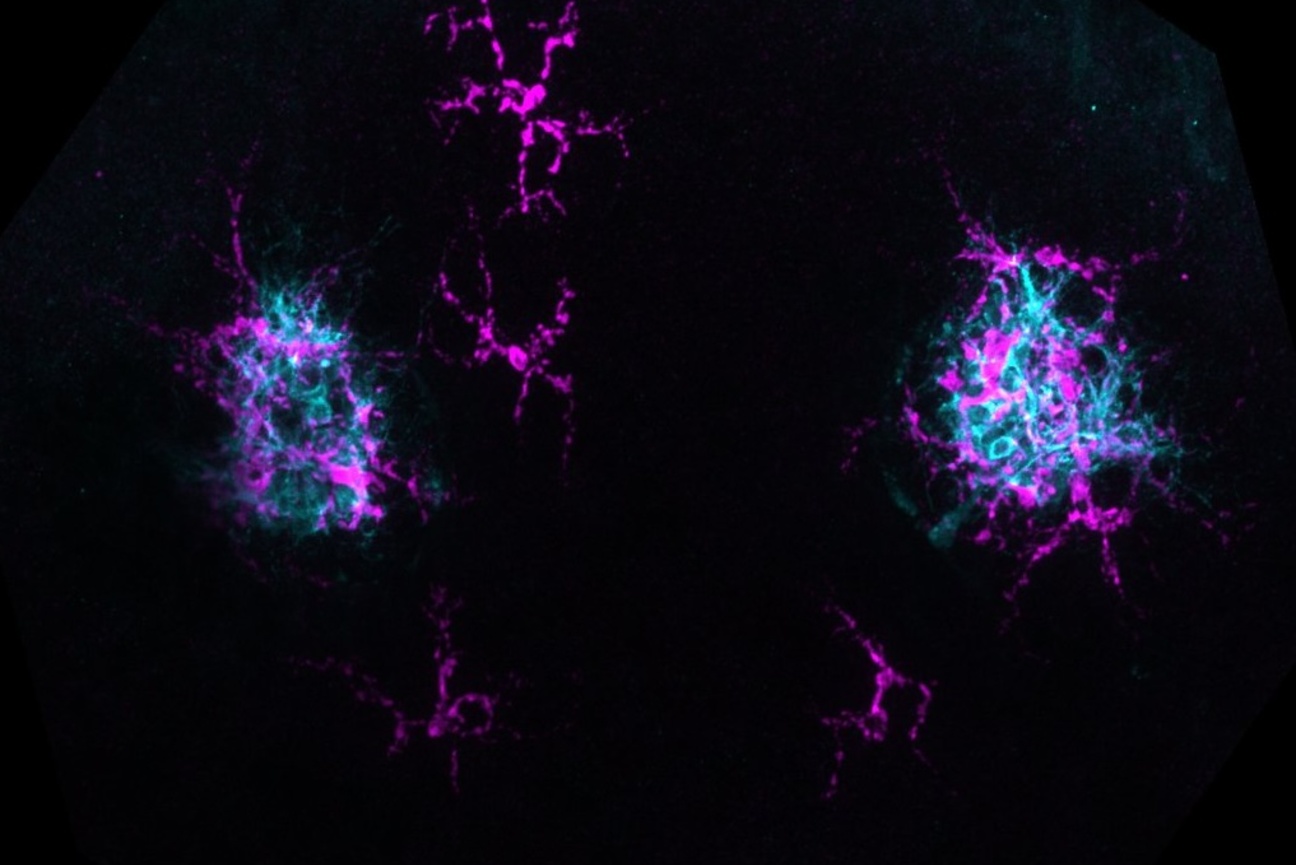
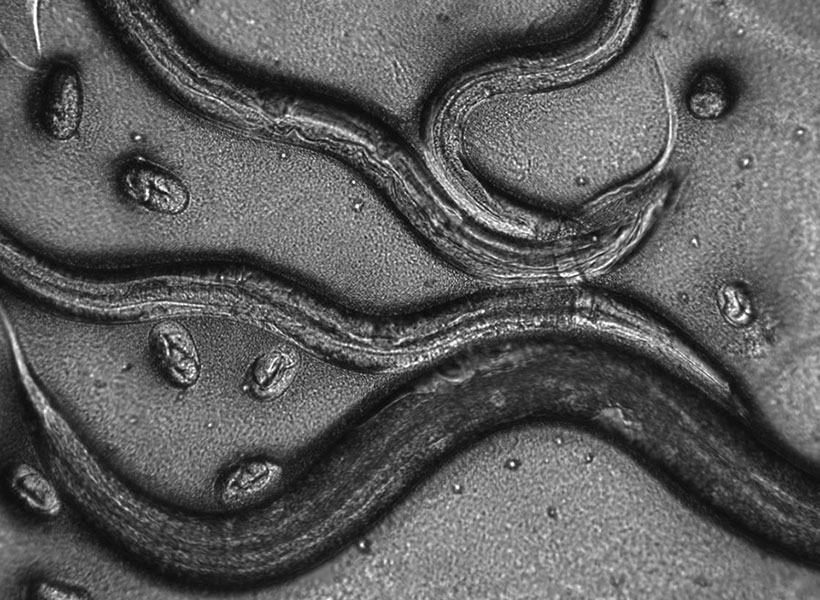
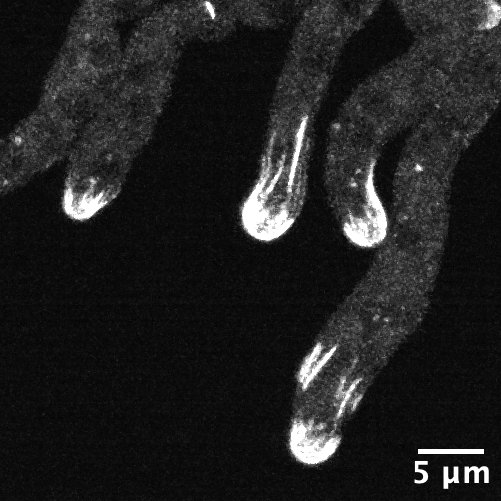
The Lew Lab works with two different specimens: a model baker’s yeast that multiplies by producing a round growth called a bud that eventually separates into a separate, daughter cell; and Aureobasidium pullulans, which is unusual because it can create multiple buds at the same time, and can also spread in large networks of branching, rootlike growths called hyphae. A. pullulans is an emerging model system, meaning that researchers are still defining what normal growth and behavior is for the fungus, like how it senses and responds to obstacles, and how resources and molecular machinery are allocated to its branching structures.
“I’m really interested in all the diversity of biology that we don’t get to study if we’re only focused on the model species,” says Clara Fikry, a graduate student in the Lew Lab and Gonzalez’s mentor for the summer.
On the mentoring side, Fikry learned how to balance providing a rigorous workload while not overwhelming her mentee with information.
“Science should be fun,” Fikry says. “The goal of this isn’t to produce as much data as possible; it’s to learn what the process of science is like.”
Although her day-to-day work was with Fikry, Gonzalez also received guidance from Daniel Lew himself. For example, his advice was invaluable for honing a draft of her research statement for potential graduate school applications, which she’d previously written as part of a class assignment.
“It was an assignment where I needed to hit a page count, and he pointed out that I kind of wrote the same thing three times in the first paragraph,” she shares with a laugh. He helped her understand that “when you’re writing something professionally, you want your writing to be concise and understandable to a broad spectrum of readers.”
Life in the cohort
The BSG-MSRP-Bio program gives undergraduate students a taste of what the day-to-day life of graduate school might feel like, from balancing one’s workload and reading research papers to learning new techniques and troubleshooting when experiments don’t go as planned. Gonzalez recalls that the application process felt very “adult” and “professional” because she was responsible for reaching out to the faculty member of the lab she was interested in on her own behalf, rather than going through a program intermediary.
Gonzalez is one of just three students from Massachusetts participating in the program this year—the program draws students from across the globe to study at MIT.
Every student also arrives with different levels of experience, from Gonzalez, who can only work in a lab during the school year about once a week, to Calo Lab student Adriana Camacho-Badillo, who is in her third consecutive summer in the program, and continuing work on a project she began last year.
“We’re all different levels of novice, and we’re coming together, and we’re all really excited about research,” Gonzalez says.
Gonzalez is a Gould Fellow, supported at MIT through the generous donations of Mike Gould and Sara Moss. The program funding was initiated in 2015 to honor the memory of Gould’s parents, Bernard S. and Sophie G. Gould. Gould and Moss take the time to come to campus and meet the students they’re supporting every year.
“You don’t often get to meet the person that’s helping you,” Gonzalez said. “They were so warm and welcoming, and at the end, when they were giving everyone a nice, firm handshake, Mike Gould said, ‘Make sure you keep going. Don’t give up,’ which was so sweet.”
Gonzalez is also supported by Cedar Tree, a Boston-based family foundation that primarily funds local environmental initiatives. In the interest of building a pipeline for future scientists with potential interest in the environmental sciences and beyond, Cedar Tree recently established a grant program for local high school and undergraduate students pursuing STEM research and training opportunities.
Preparing for the future
The BSG-MSRP-Bio program culminates with a lively poster session where students present their summer projects to the MIT community—the first time some students are presenting their data to the public in that format.
Although the program is aimed at students who foresee a career in academia, the majority of students who participate are uncertain about the specific field, organism, or process they’ll eventually want to study during a PhD program. For Gonzalez, the program has helped her feel more prepared for the potential rigors of academic research.
“I think the hardest thing about this program is convincing yourself to apply,” she says. “Don’t let that hinder you from exploring opportunities that may seem out of reach.”