Over six years of operation, pre-college outreach programs administered by Mandana Sassanfar, Senior Lecturer and Director of Diversity and Outreach, have placed seven exceptional pre-college students, often from underserved or underrepresented backgrounds, with research groups in The Picower Institute.
David Orenstein | The Picower Institute for Learning and Memory
During the pandemic, when many classes delivered online could barely hold students’ attention, Presley Simelus became captivated by the subject of biology thanks to their boundless curiosity and their uncommonly engaging teacher at Prospect Hill Academy Charter School in Cambridge. Meanwhile for Eli Hanechak, the science bug must have bit her very early. She’s wanted to be a doctor for as long as she can remember and in fifth grade built a model of a space station the size of a car out of duct tape, cardboard and broomsticks.
Not every teenager is expected to want to spend their summer breaks exploring science at a bench in an MIT lab, but each year students like Simelus and Hanechak, who have a distinct passion for research, can bring that to The Picower Institute and other research entities around MIT. Over six years of operation, pre-college outreach programs administered by Mandana Sassanfar, Director of Diversity and Outreach, have placed seven exceptional pre-college students, often from underserved or underrepresented backgrounds, with research groups in The Picower Institute. Despite their relative lack of experience compared to the technicians, graduate students, postdocs and professors around them, the students typically thrive.
“Eli has been a wonderful addition to our lab for the summer,” said Kendyll Burnell, the graduate student in the lab of Professor Elly Nedivi who has been working closely with Hanechak. “She is a hard worker, has caught on to techniques quickly, and is constantly asking excellent questions about science and doing research.”
Simelus, too, has been not only learning but also contributing, said their summer host, Yire Jeong, a postdoc in the lab of Associate Professor Gloria Choi.
“Presley has been amazing in our lab, and I was impressed by Presley’s eagerness to learn so much about neuroscience,” Jeong said. “Even when facing technical difficulties, Presley diligently worked to overcome them and achieved meaningful results.”
‘Dive into it’
Simelus, who hails from Everett, Mass., and will be enrolling in Swarthmore College this fall to study biochemistry, first came to MIT through the Leah Knox Scholars Program. Friends who’d been in the program before encouraged them to apply and they got in. During five weeks last summer Simelus and their cohort of fellow Leah Knox high-schoolers had the geeky pleasure of extracting bacteria out of the Charles River and performing a battery of tests to genetically characterize the novel organisms they found. Sassanfar noted that Simelus did the lab work exceptionally well, which is something she looks for when determining whom she might invite back the next summer to do research in an MIT Brain and Cognitive Sciences or Biology lab.
This spring when it came time for Simelus to decide where they might like to take that opportunity, they chose the Choi lab, which studies how the central nervous systems and immune systems interact, sometimes with consequences relevant to disorders including autism. Those keywords intrigued Simelus but really they made the choice because of the potential to learn something entirely new.
“It was all this stuff I just simply wasn’t familiar with and I wanted to learn more about it,” Simelus said. “With Gloria’s lab I was truly mystified and I wanted to dive into it. That’s the reason I chose it.”
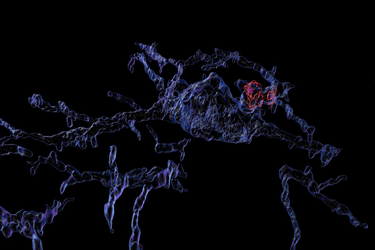
This summer Simelus has been working with Jeong on a study of how brain cell activity differs when mice are sick vs. when they are well. The project has involved imaging neurons in the brain to detect telltale signs of recent activation, expression of a protein called c-fos. Learning about neuroscience and gaining skills like preparing, staining and imaging tissue have been a very fulfilling outcome of the internship, Simelus said.
“I truly have learned so much about neuroscience,” they said. “I feel like the field, anything related to the brain or neuroscience, is always under this sort of veil and nobody really knows what’s going on. But I feel like my time at the Choi lab has really allowed me to see what neuroscience is about. It’s taught be more about the brain itself and also more about different biology techniques and skills I might need.”
Now the only problem, Simelus said, is that there are even more things to be deeply curious about. Simelus feels committed to harnessing the life sciences in some way in the future to sustain human life and experience. And as someone who not only plays the viola but also composes, they’ve begun thinking more about how the brain responds to music.
There will no doubt be many chances to continue exploring these interests at Swarthmore, but during the summer at MIT, Simelus said they’ve expanded their horizons while still hanging out with friends, some of whom have been working in other nearby labs.
“I don’t think I would have changed my summer,” Simelus said.
‘The perfect opportunity’
Hanechak lives in the tiny Western Massachusetts town of Russell (population: 1,643) and commutes 45 minutes to Pope Francis Preparatory School in Springfield, where she is a rising senior.
In her freshman year at a different school, she yearned for an extra challenge so she got involved in science fair. Interested in medicine, but eager for a project in which she could make a difference without having clinical credentials, she chose to work on reducing pollution by developing a microbe-derived enzyme that could biodegrade plastics. She had read about such enzymes in the research literature and learned that they don’t work as well as engineers have hoped. In successive years she has scrounged lab space and general supervision in labs at Westfield State University and UMass Amherst to create and screen beneficial mutations in the enzyme and to synthesize structures that might help the enzyme work better. The enzyme she presented at the International Science and Engineering Fair last year can degrade plastics in 24 hours.
Sasssanfar, who also directs the Massachusetts Junior Academy of Science (MassJAS), learned of Hanechak’s award-winning science fair presentation and invited her to present at the MassJAS symposium, held at MIT last October. Hanechak did so well, Sassanfar said, she earned a spot present at the American Junior Academy of Science meeting (adjacent to the American Association for the Advancement of Science Annual Meeting) in Denver in February. She also earned Sassanfar’s invitation to join a lab this summer at MIT.
Hanechak has long had an MIT pennant on her wall at home and has admired MIT as a place where regardless of one’s background, if one has a passion for science and technology, that’s what matters.
“No one in my family has gone to college and no one has been involved in a science-related career of any kind,” she said. “One of the reasons MIT has always stood out to me is that there are especially great minds here, but they didn’t all come from established families or super prestigious backgrounds or anything like that. They kind of just were able to make their own way.”
Moreover, the chance to come to MIT to learn about the brain in the Nedivi lab seemed like a great step to take toward that longer-term goal of medicine.
“It seemed like the perfect opportunity to start transitioning into what I want my career to look like and to get some experience doing neuroscience research,” Hanechak said. “I’m very glad I’m able to have this summer experience, like learning the techniques. When I go into my college major of neuroscience, I will have a good background of what I’m doing, besides just my environmental research.”
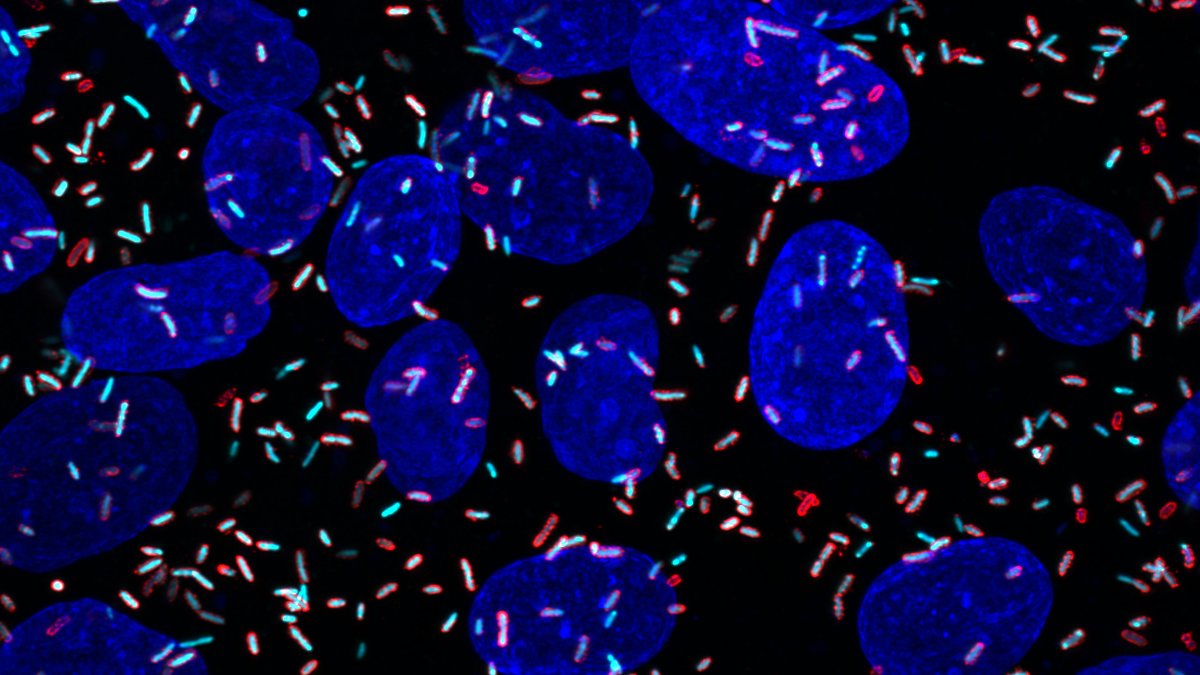
With Burnell, Hanechak is working on finding a DNA promoter specific for a rare but interesting kind of neuron in the visual cortex, where the brain processes what the eyes see. Finding this genetic signature would allow the lab to label these cells and image them under the microscope, so that they could see how the cells contribute to visual processing.
Hanechak acknowledged she was anxious at first about joining a bigger lab with scientists who have much more experience.
“But my entire summer has been incredibly gratifying and exciting—just being able to work in Cambridge, and live in this area, and experience city life, and then also be in a lab environment where it’s so collaborative and everyone’s very friendly,” she said.
For many teens, summer provides a chance to do what they want to do. Simelus and Hanechak chose the opportunity to explore the brain at The Picower Institute and have made the most of it.