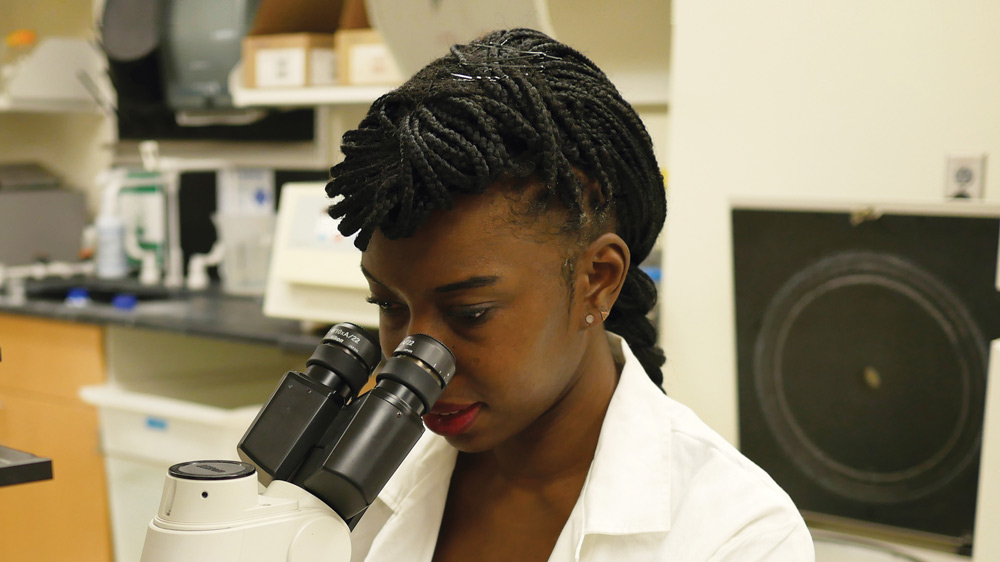
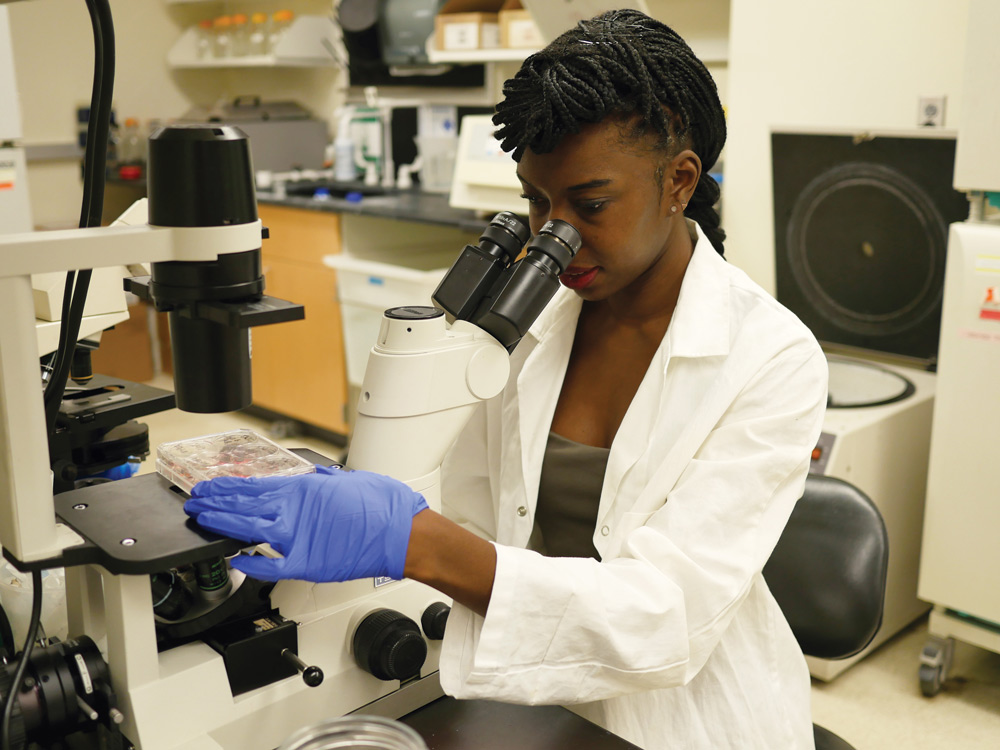
Elizabeth Li ’18 has worked in three cancer-related labs over the past six years, and one day intends to start her own.
Raleigh McElvery
March 6, 2018
Senior Elizabeth Li recreates miniature organs — lungs and intestines — outside the body. She does so in order to study cancer progression in both environments, and over the past six years has worked in three separate cancer-focused labs: two at MIT and another beginning her junior year of high school. One day, she aims to run her own.
“I’ve been into math and science ever since I was little,” she explains, “but in third grade I met a friend who was pretty important to me. She was diagnosed with a very malignant form of brain cancer and ended up dying from it. From that point on — even though I was still very young — I knew I wanted to do cancer research.”
In 9th grade, Li began at the School for Science and Math at Vanderbilt, a joint program between the university and Metro Nashville Public Schools. “I got to skip school once a week to learn research techniques, and had the opportunity to join Andries Zijlstra’s lab my junior year,” she recalls. “I’m actually still part of that group, and I’ve been working on the same project related to cancer metastasis for six years now.”
When it came time to select a university, Li was torn between Vanderbilt — where she was already performing research — and MIT, which she describes as “the place of opportunity.” She was ultimately swayed by MIT’s vast array of research areas, and fully sold after an overnight to preview the undergraduate culture.
Li opted for Course 7 in order to continue doing cancer research, and joined Omer Yilmaz’s lab in 2015 to investigate intestinal tumorigenesis. Here, she spent most of her time doing organoid work, studying the progression of colorectal cancer in miniaturized and simplified versions of the intestine. Li removed individual intestinal stem cells — or sometimes the entire “crypts” in which they reside — and grew them inside a 3D gel. This environment allowed the cells to differentiate and interact as they would in the colon, rather than growing on a flat, plastic dish.
Li and other members of the Yilmaz lab watched these cells multiply, observing their shape and the regeneration process. Li’s method of assessment varied depending on the research question: on some days, she stained the cells for proliferation markers, and on others she exposed them to different metabolites or drugs to see how the cells responded.
“On a typical day, I would come in during the morning between classes, and again in the afternoons and evenings,” she says. “The experiments differed, but we tended to do a lot of genetic manipulation. We’d make plasmids, CRISPR-Cas9 knockouts, or test for gene and protein expression using qPCR and Western blots.”
After two years, Li’s primary mentor finished her postdoctoral training, and Li transitioned to Jackie Lees’ lab at the beginning of her senior year. Li is now working with a fellow undergraduate on an independent project, centered on the enzyme protein arginine methyltransferase 5 (PRMT5).
PRMT5 catalyzes the transfer of methyl groups to the amino acid arginine in certain proteins, modifying their function. The enzyme also plays a key role in regulating gene splicing, the process by which segments of pre-mRNA are removed — changing the genetic code so that multiple genes can be encoded by the same initial transcript.
The Lees lab is interested in PRMT5 because it affects glioblastoma formation, the most common form of adult brain malignancy. As Li explains, when PRMT5 expression increases, so does tumor formation. Since there are still relatively few therapeutic options to treat glioblastomas, she’s hoping to use small molecules to inhibit PRMT5 expression and thus impede tumor initiation and progression.
“We’re considering using nanoparticles to deliver them,” she says, “and in doing so, hoping to gain a better understanding of how PRMT5 inhibition might impact cancer progression and tumorigenesis.”
Li is testing one small molecule PRMT5 inhibitor in lung organoids and several 2D cell lines — determining how sensitive the cells are and if the organoids will form, to gauge whether a tumor would still develop in the presence of the drug. “Depending on when you add the drug, you can test different aspects of tumorigenesis,” she explains.
She’s also split the past four years between the Biology Undergraduate Student Association (of which she was faculty liaison, outreach chair, and then co-president), the MIT Pre-Medical Society, MIT Lion Dance, Asian Dance Team, Wind Ensemble, and Improv-a-Do! She’s also heavily involved in DynaMIT, which organizes an annual, week-long science program for economically disadvantaged middle school students.
“There are a lot of extracurriculars to do,” Li admits. “But it’s pretty easy to get involved in the MIT community and still stay on top of your coursework, if you keep it to four or five classes per semester. It’s worked out for me so far — I’m still alive and happy and have time for eating, sleeping, and friends.”
As Li applies to MD-PhD programs, she hopes one day to practice medicine (perhaps pediatric oncology) while running her own lab.
“My advice for incoming MIT undergrads would be to remember to have fun,” she says. “You only have four years, so take advantage of your time here: hang out with your friends, take the classes you want to take, and do things that you enjoy. Hopefully most of those activities will be one and the same.”