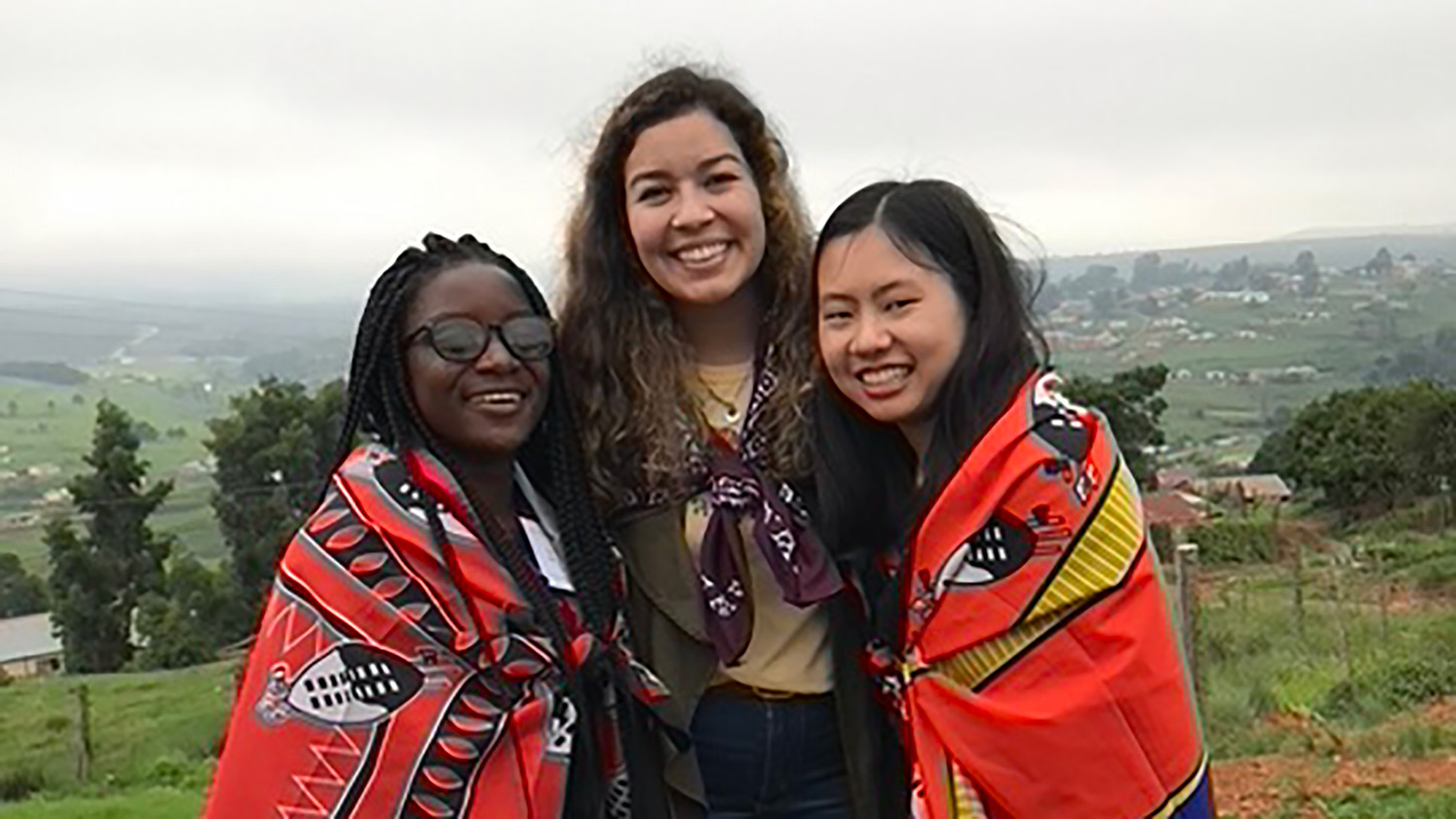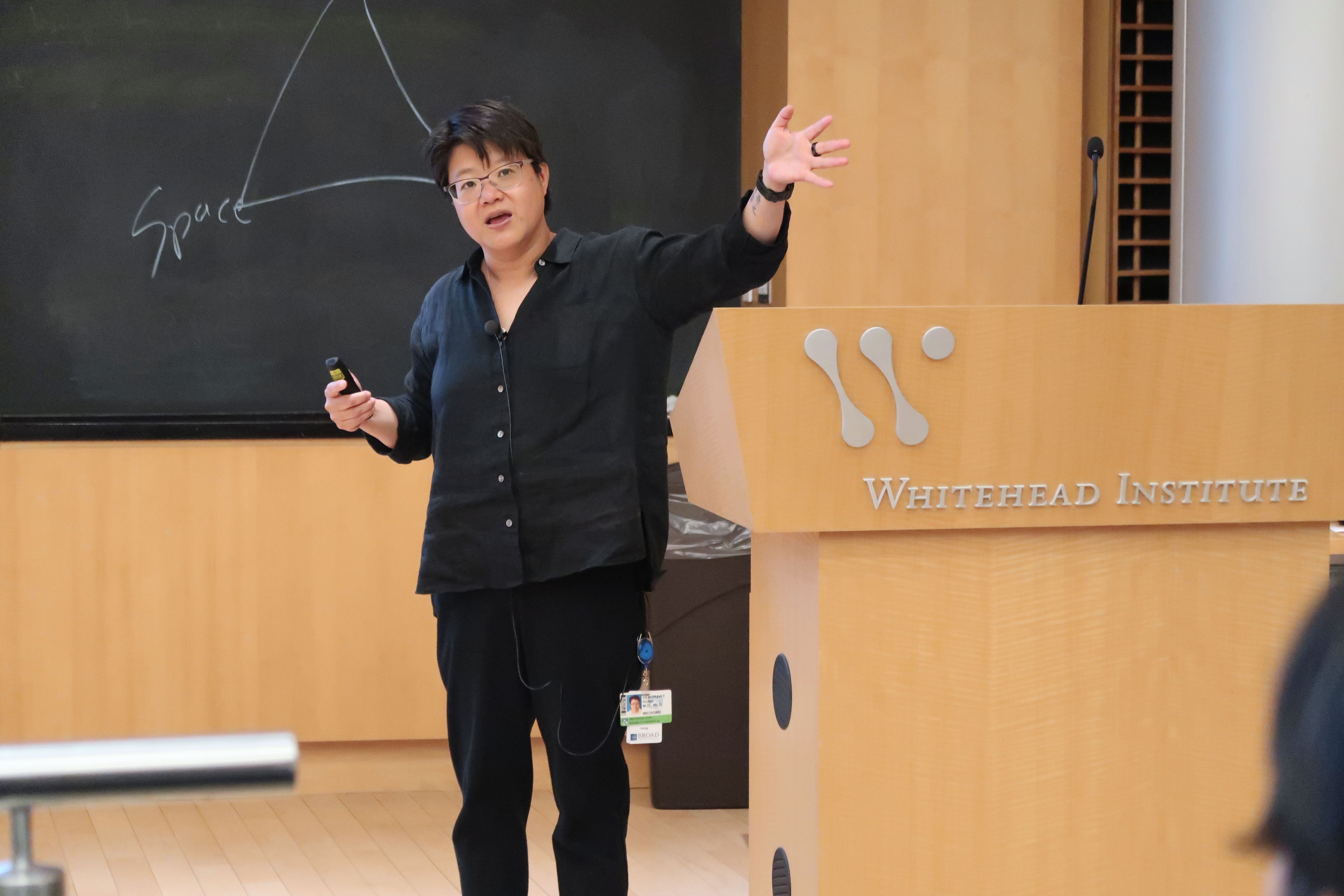
Emma H. Yee, Steven S. Cheng, Grant A. Knappe, and Christine A. Moomau | MIT Science Policy Review
August 25, 2020
Article Summary
Antibiotics are a vital component of global health. By killing or inhibiting the growth of bacteria, antibiotics treat infections like pneumonia, staph, and tuberculosis.By preventing infections, they enable major medical procedures such as surgeries and chemotherapy. However,bacteria are becoming increasingly resistant to current antibiotics, causing an estimated 34,000 deaths annually in the US. Left unchecked, antibiotic resistance will have major public health consequences, causing over 5 million deaths each year by 2050. Major causes of this crisis are the misuse of existing antibiotics and the slow development of new antibiotics. To incentivize responsible use, governments and institutions are initiating education programs, mandating comprehensive hospital antibiotic stewardship programs, and funding the development of rapid diagnostics. To bring new antibiotic drugs to market, the US government and other non-governmental organizations are funding scientific research toward antibiotic development.Additional incentives are being pursued to improve the commercial viability of antibiotic development and protect drug developers from the unique challenges of the antibiotic market. With diligent efforts to improve responsible use and encourage novel antibiotic drug discovery, we can decrease the global disease burden, save money, and save lives.
Antibiotics are drugs that kill or inhibit the growth of bacteria, and we have them to thank for the 25-year increase in American life expectancy in the last century[1,2]. In 1900, the three leading causes of death were bacterial infections: pneumonia, tuberculosis, and diarrhea/enteritis[3]. Penicillin, the first antibiotic, was discovered in 1928. But it was not until World War II, when wounded soldiers were more likely to die from infections than the injuries themselves, that governments realized penicillin’s life-saving potential[4]. The US government began developing and mass-producing penicillin through unprecedented public, private, and international collaborations, prompting a new era of antibiotics. Antibiotics are now used to treat a myriad of common infections like strep throat, meningitis, tuberculosis, tetanus, urinary tract infections, and food poisoning. They also enable medical procedures that otherwise create a high risk of infection, such as invasive surgery, organ transplantation, and chemotherapy[5]. However, antibiotics are not “one size fits all”; certain types of antibiotics are only effective against certain kinds of bacteria, and all antibiotics are ineffective against viruses[6].
Antibiotics kill or inhibit bacterial growth via various mechanisms of action; they might attack the protective bacterial cell wall, interfere with bacterial reproduction, or interrupt production of molecules necessary for the bacteria’s survival[7]. However, bacteria reproduce and evolve rapidly, changing over time to resist an antibiotic’s destructive mechanism of action. In fact, the more we use antibiotics, the faster bacteria evolve to resist those antibiotics. As bacteria reproduce, random DNA mutations will occur. Most random mutations have no effect on the bacteria, but sometimes a mutation will give the bacteria a special ability to resist an antibiotic—for instance, the mutation may change the cellular target of the antibiotic, or allow the bacteria to pump the drug out of the cell. When an antibiotic is used on bacteria, most of the population will die, but if any of the bacteria have one of these resistance-conferring mutations, they will survive and continue to reproduce, until the entire population is resistant[5]. The use of antibiotics therefore creates environments where bacteria with antibiotic resistance mutations are more likely to survive and reproduce, while susceptible bacteria are gradually killed off.

This means that, over time, the bacteria that cause infections in humans are more and more likely to be resistant to common antibiotics. It is important to note that bacteria develop antibiotic resistance–not people. But when people use lots of antibiotics, they change bacterial populations such that more and more bacteria are resistant to those antibiotic drugs. This illustrates the double-edged sword nature of antibiotic use: antibiotics are immensely valuable for combating countless infections and enabling medical procedures, but the more we use them, the less valuable they become.
Today, antibiotic resistance is accelerating at alarming rates. The Centers for Disease Control and Prevention (CDC) estimates there are 3 million antibiotic resistant infections in the US every year, causing at least 34,000 deaths[5]. Globally, at least 700,000 deaths occur due to resistant infections, most of which are bacterial; the actual number is likely higher due to poor reporting and surveillance[8]. The prospect of widespread antibiotic resistance threatens to bring society into a post-antibiotic age where infections are more expensive and difficult to treat. This is a threat to not only public health but also the economic stability of the healthcare system[9] and national security[10].

Misuse and Overuse of Antibiotics
Overuse of antibiotics is a major contributor to the rapid proliferation of antibiotic resistant infections. It is estimated that US doctors’ offices and emergency departments prescribe about 47 million unnecessary antibiotic courses annually, amounting to 30% of all antibiotic prescriptions[12]. Many studies show that even when illnesses do require antibiotics, prescribed time courses are significantly longer than national guidelines[13, 14].
Rapid Diagnostics and Antibiotic Prescriptions: A major cause of ubiquitous antibiotic overuse is a lack of rapid methods for diagnosing infections. Physicians rely on tests that usually take days to weeks to identify if an infection is bacterial and, if so, which antibiotics will be most effective. Waiting this long can be harmful or even fatal for patients[15]. Therefore, physicians usually prescribe broadly effective antibiotics while knowing little about the nature of the infection[15]. This can save lives, but if the infection is caused by a virus or resistant bacteria, the antibiotics will not treat the illness and will give resistant strains a chance to further multiply, leaving patients susceptible to additional infections.
With growing awareness in the last 5-10 years that appropriate antibiotic use is difficult with current diagnostics, the CDC, the National Institute of Allergy and Infectious Diseases (NIAID), and the Biomedical Advanced Research and Development Authority (BARDA) have collectively awarded hundreds of millions of dollars to state health departments, businesses, and universities to develop rapid diagnostics[16]. BARDA and NIAID also organized a $20 million prize, the Antimicrobial Resistance Diagnostic Challenge[17], and fund the global non-profit, CARB-X, which has invested $82.5 million in 55 projects worldwide for antibiotic resistance research, including diagnostics[18]. This surge in resources and funding has increased rapid diagnostic development. For example, the NIAID funded development of BioFire’s FilmArray[19], which is now an FDA-cleared diagnostic test available for purchase in the US[20]. In just an hour, it tests patient samples for several common types of bacteria, viruses, and yeast, including antibiotic resistant ones[21].
However, new diagnostic technologies have limited effectiveness when they fail to meet practical cost and resource requirements. Cepheid’s GeneXpert MTB/RIF test, for example, can diagnose tuberculosis infection and determine resistance to rifampicin, a common antibiotic for tuberculosis, in 2 hours[22]. Unfortunately, it has not been used as widely as initially expected[23], mainly because the equipment costs $17,000, not counting training and set-up costs[24]. This illustrates another major shortcoming of current diagnostic technologies: high healthcare infrastructure and cost requirements that render them inaccessible to many people.
Widespread access to rapid diagnostics is not just about fairness, it’s a necessity. Antibiotic resistance will remain a problem in the US as long as it is a problem anywhere in the country or the world due to inevitable intra- and international bacterial transmission. Many recently developed rapid diagnostics cost $100-$250 per test[25, 26]. These diagnostic innovations are promising and valuable in filling part of the gap in rapid diagnostics, but their benefits will not be felt by the majority of global hospitals and patients that cannot afford or support high cost, high tech diagnostic investments. Increasing institutional funding in the last 10 years has resulted in new rapid diagnostics for identifying and characterizing infections, a potential step towards reducing antibiotic misuse and subsequent development of antibiotic resistance. However, ensuring accessibility of technological improvements is essential in combating antibiotic resistance.
Prescribing Practices: Updating prescription standards and educating healthcare workers and patients on responsible antibiotic use is another key step in reducing antibiotic overuse. In the US, patients are often prescribed antibiotics for far longer than necessary. Two recent studies found that 70% of patients with sinus infections and 70% of adults hospitalized with pneumonia were given antibiotics for 3 or more days longer than recommended[13, 14]. Oftentimes, this stems from an out-of-date belief that longer is better in terms of preventing the development and spread of resistant bacteria. In fact, the opposite is true. Shorter courses of antibiotics lower the selective pressure for development of resistance. This was illustrated in a study of pediatric antibiotic use[27], where children prescribed 5 days of amoxicillin for the treatment of respiratory infections were less likely to carry antibiotic resistant Streptococcus pneumoniae in their nasal passage than their peers who were treated for 10 days. These children were also found to be less likely to transmit resistant bacteria to others.
In many cases, common antibiotic treatments can be shortened without affecting the outcome. A trial of pneumonia patients found that the standard 8-day course of amoxicillin can be shortened to just 3 days with equal symptom relief and fewer side effects[28]. Similarly, treatment of ventilator-associated pneumonia can be effectively shortened from 14 to 8 days[29]. In some cases, shortened antibiotic courses have actually improved patient outcomes. A reduced course for urinary tract infections from 14 days to 7 days is not only effective, it also prevents post-treatment yeast infections[30].
As scientists and clinicians become more aware of the dangers of resistance, more studies are being conducted to determine the minimum amount of antibiotic required to adequately treat infections. The Infectious Diseases Society of America has also updated their Clinical Practice Guidelines to reflect findings that shorter treatment schedules are often just as effective, are easier to comply with, and reduce development and spread of resistant bacteria[31]. Performing
minimum effective antibiotic treatment trials is costly in the short term, but necessary to safely revise guidelines and save on long-term healthcare costs.
Public misunderstanding and misinformation regarding antibiotics also contribute to their overprescription. In many clinical settings where antibiotics are not necessary, patients may believe antibiotics are the most effective treatment and push their doctors to inappropriately prescribe them. For example, patients often seek antibiotics for viral respiratory illnesses (i.e. cold and flu), despite antibiotics being ineffective against viral infections[5]. It has been demonstrated that patient expectation of antibiotics or physician perception of this desire have a significant influence on antibiotic prescription[32–34].
Table 1: Antibiotic overuse is caused largely by shortcomings in diagnostic technologies and prescribing practices, but there are many possible ways to address these challenges.

Revitalizing the Antibiotic Pipeline
While it is important that existing antibiotics are prescribed cautiously and used responsibly, all antibiotics inevitably encounter resistance[38]. Consequently, continuously developing antibiotics with novel mechanisms of action—the method that an antibiotic uses to kill bacteria—that circumvent existing resistances will remain essential. However, developing these new drugs is costly; it can take well over a decade and cost more than $2 billion, with a 90% failure rate looming over the project[38]. Clinical trials, which require large, diverse populations to demonstrate evidence of drug superiority, account for 65% of the risk-adjusted cost for developing antibiotics[15]. The difficulty of antibiotic drug development is illustrated by the 2019 FDA approval of lefamulin, which marked the first approval of an IV/orally-administered antibiotic with a novel mechanism of action in two decades[39]. Scientific challenges inhibit discovery significantly. The immediately apparent antibiotic candidates have been developed, and discovering antibiotics with new mechanisms of action is challenging. It is now thought that any new, effective antibiotics will need multiple capabilities for killing bacteria, making their discovery more complex[3]. Emerging approaches in antibiotic discovery such as deep learning algorithms are promising technologies to solve these scientific challenges, but are far from bringing new antibiotics to patients[40].
In addition to scientific obstacles, the economics of antibiotic development have reduced innovation and output. The free market is failing to meet society’s antibiotic needs via multiple pathways[41]. Traditional sales-based models, in which revenue is directly proportional to the volume of sales, are antagonistic towards society’s goal of sustainable antibiotic use[2]. Evidence of the current system’s failure is the drastic decrease in antibiotic research programs[3] and the sparse output of new [2]. To address these challenges, policymakers are crucial actors; they can facilitate fertile economic conditions using a combination of 1) “push” policies to galvanize antibiotic discovery and development efforts and 2) “pull” policies to create profitable
economic conditions, incentivizing industry to work in this area. Simultaneously, these policies must be supplemented by sufficient regulations to ensure sustainable and equitable usage, broadly maximizing overall societal benefits.
Push Policies: Push policies drive companies to conduct antibiotic research and clinical trials[42] by providing monetary resources to antibiotic developers. Push policies are realized via grants and pipeline coordinators. Government grants allow both academia and industry to investigate antibiotic candidates and conduct clinical trials. Pipeline coordinators are agencies that ensure governmental funding is distributed efficiently across development stages. Coordinators are essential to ensuring equitable funding distribution across antibiotic candidates and identifying gaps and needs in the antibiotic pipeline from basic research through production. These vehicles have broad precedents and have demonstrated effectiveness at stimulating early stage scientific research. Current estimates show $550 million is spent annually on push spending, though some recommendations show that this number should be $800 million to fully meet the demand for antibiotic research[42]. However, push policies and spending do not completely address the major economic issues.

Pull Policies: The primary goal of push policies is to jump start research and development in antibiotic discovery, but issues remain with the current market structure for antibiotics. This is illustrated by the fact that companies are failing after bringing important antibiotics to market. For instance, the biopharmaceutical company Achaogen successfully developed the antibiotic plazomicin in 2018, but filed for bankruptcy the following year due to insufficient profits from plazomicin[43]. Why would a company that successfully brings a new antibiotic to market fail? Antibiotics are generally prescribed for short periods of time (usually under two weeks), modern health policies support reducing or delaying the use of new antibiotics, and the market lifetime of antibiotics is reduced due to the inevitable development of resistance[44].Overall, these realities minimize sales of the new antibiotic and thus the profits of the developing company. In response, policymakers have proposed pull policies to de-link the sales of the new antibiotic to the economic reward given to the developers, improving the economic viability of developing new antibiotics. These pull policies are supported by the Infectious Diseases Society of America[45]. By de-linking sales from economic reward, the revenue from a new antibiotic is not purely based on the sales volume of that antibiotic. For example, a market entry reward (MER) — a large monetary sum given to developers of novel antibiotics upon successful drug approval — can be used to partially or fully de-link the number of sales from the economic reward. Multiple groups, such as the Boston Consulting Group, have estimated that a $1 billion MER per antibiotic is sufficient, suggesting that this award amount would lead to twenty novel antibiotics for society over the next three decades[42, 46].
An important supplement to any MER policy is the antibiotic susceptibility bonus (ASB)[47]. The ASB rewards companies that develop antibiotics that are effective over long periods of time. As an antibiotic remains effective against target bacteria, companies receive monetary awards. This policy helps better align all stakeholders’ (companies, patients, hospitals, insurance networks) interests towards generating and maintaining effective antibiotics. Companies will no longer have an incentive to oversell antibiotics, as they will receive more money the longer their drug is effective. This supplemental policy could safeguard MERs against abuse, and incentivize the development of antibiotics that act in society’s best interest: to develop effective treatments for long periods of time.
Another potential pull policy is the long-term supply continuity model (LSCM)[42], which addresses how companies respond once market exclusivity for a drug ends due to patent expiration. Suppliers may respond to loss of market exclusivity by either manufacturing fewer units in the case of a modest market or by increasing sales through marketing and promotion. Both actions are detrimental to public health in the case of an antibiotic, either promoting antibiotic overuse or making it harder for people who need the antibiotic to get it. The LSCM addresses this by having a country or group of countries make an agreement with manufacturers to produce a predetermined amount of the respective antibiotic for a specified price. This model to generate a predictable supply of an antibiotic acts as a pull mechanism by making the market for novel, essential antibiotics more sustainable for manufacturers.
Pull policies also have some downsides. For one, pull policies only reward successful antibiotic discovery campaigns; the inherent risk in developing these drugs may still dissuade companies. Also, while push policies have been validated with real world results, pull policies have not been evaluated as extensively. To encourage companies to work in this area, push policies, as well as pull policies, are needed to lower the risk of failed discovery programs. To develop the new drugs that society needs, companies need funding to start research and development and economic incentives to take the drugs to market.
Acknowledgements
We thank Erika Madrian for her input in shaping the manuscript.
Citation
Yee, E. H., Cheng, S. S., Knappe, G. A. & Moomau, C. A. Antibiotic resistance: How to prevent the next public health emergency. MIT Science Policy Review 1, 10-17 (2020).