Raleigh McElvery
July 9, 2019
Listeria monocytogenes, the food-borne bacterium responsible for listeriosis, can creep from one cell to the next, stealthily evading the immune system. This strategy of cell-to-cell spread allows them to infect many different cell types, and can spur complications like meningitis. Yet the molecular details of this spread remain a mystery.
In a paper recently published in Molecular Biology of the Cell, researchers from the MIT Department of Biology, University of California, Berkeley, and Chan Zuckerberg Biohub are beginning to piece together the elusive means by which Listeria moves from one cell to the next. This mode of transport, the scientists suggest, looks a lot like trans-endocytosis, a process that healthy, uninfected cells use to exchange organelles and various cytoplasmic components. In fact, the two processes are so similar that Listeria may be co-opting the host cell’s trans-endocytosis machinery for its own devices.
Although the particulars of trans-endocytosis are poorly understood, the process permits neighboring cells to exchange materials via membrane-bound compartments called vacuoles, which release their cargo upon reaching their final destination.
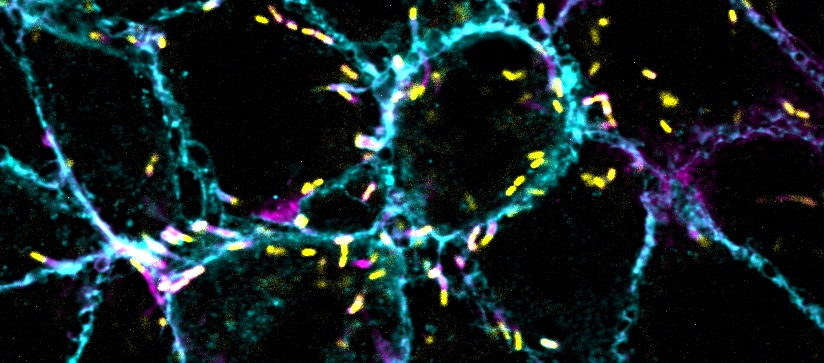
Much like trans-endocytosis, cell-to-cell spread relies on vacuoles to ferry Listeria. First, the pathogen commandeers the host cell’s own machinery to assemble a tail of proteins that allows it to rocket around inside the cell and ram against both the membrane of the host and that of the adjacent cell. The resulting protrusion is then somehow engulfed into a double-membrane vacuole, and the bacteria burst through their containment to begin the process anew in the recipient cell.
“There’s been a lot of work looking at Listeria cell-to-cell spread,” says Rebecca Lamason, the Robert A. Swanson (1969) Career Development Assistant Professor in the MIT Department of Biology and senior author on the study. “But we still don’t really understand the molecular mechanisms that allow the bacteria to manipulate the membrane to promote engulfment. Depending on what we uncover, we might also be able to apply that information to better grasp how an uninfected cell regulates trans-endocytosis.”
Lamason and her team anticipated that the same proteins implicated in trans-endocytosis would also be involved in Listeria cell-to-cell spread, which would indicate that the pathogen was appropriating these proteins for its own purposes. The researchers made a list of 115 host genes of interest, and then used an RNAi screen to identify just 22 that are critical for cell-to-cell spread.
They were excited to find that, of those 22 genes, several are also implicated in endocytosis, which suggests Listeria is using a similar strategy. These include genes encoding caveolin proteins that control membrane trafficking and remodeling, as well as another protein called PACSIN2 that interacts with caveolins to regulate protrusion engulfment.
Now that the researchers have pinpointed these key proteins, the next step is to determine how they work together in order to promote cell-to-cell spread — especially since the protrusions created by Listeria are much larger than those required for trans-endocytosis.
“As we drill down even deeper into the molecular mechanisms, it will be interesting to see where trans-endocytosis and cell-to-cell spread differ, and where they are similar,” Lamason says. “Our hope is that investigating the mechanisms of bacterial spread will reveal fundamental insights into host intercellular communication.”
Citation:
“RNAi screen reveals a role for PACSIN2 and caveolins during bacterial cell-to-cell spread”
Molecular Biology of the Cell, online June 26, 2019, DOI: 10.1091/mbc.E19-04-0197
Allen G. Sanderlin, Cassandra Vondrak, Arianna J. Scricco, Indro Fedrigo, Vida Ahyong, and Rebecca L. Lamason