Research Area: Biochemistry, Biophysics, and Structural Biology
Researchers have devised a faster, more efficient way to design custom peptides and perturb protein-protein interactions.
Raleigh McElvery | Department of Biology
February 15, 2019
One way to probe intricate biological systems is to block their components from interacting and see what happens. This method allows researchers to better understand cellular processes and functions, augmenting everyday laboratory experiments, diagnostic assays, and therapeutic interventions. As a result, reagents that impede interactions between proteins are in high demand. But before scientists can rapidly generate their own custom molecules capable of doing so, they must first parse the complicated relationship between sequence and structure.
Small molecules can enter cells easily, but the interface where two proteins bind to one another is often too large or lacks the tiny cavities required for these molecules to target. Antibodies and nanobodies bind to longer stretches of protein, which makes them better suited to hinder protein-protein interactions, but their large size and complex structure render them difficult to deliver and unstable in the cytoplasm. By contrast, short stretches of amino acids, known as peptides, are large enough to bind long stretches of protein while still being small enough to enter cells.
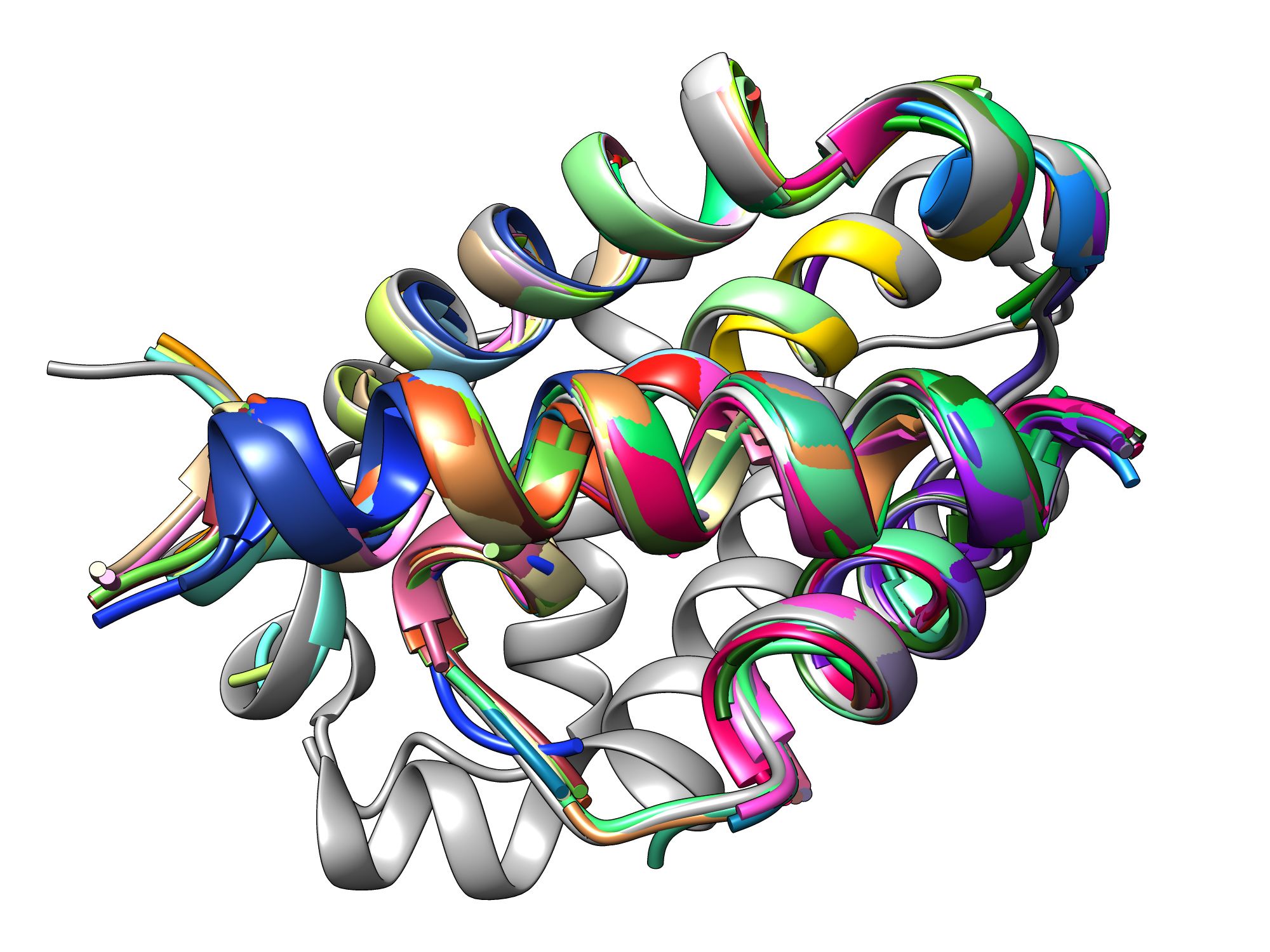
The Keating lab at the MIT Department of Biology is hard at work developing ways to quickly design peptides that can disrupt protein-protein interactions involving Bcl-2 proteins, which promote cancer growth. Their most recent approach utilizes a computer program called dTERMen, developed by Keating lab alumnus, Gevorg Grigoryan PhD ’07, currently an associate professor of computer science and adjunct associate professor of biological sciences and chemistry at Dartmouth College. Researchers simply feed the program their desired structures, and it spits out amino acid sequences for peptides capable of disrupting specific protein-protein interactions.
“It’s such a simple approach to use,” says Keating, an MIT professor of biology and senior author on the study. “In theory, you could put in any structure and solve for a sequence. In our study, the program came up with new sequence combinations that aren’t like anything found in nature — it deduced a completely unique way to solve the problem. It’s exciting to be uncovering new territories of the sequence universe.”
Former postdoc Vincent Frappier and Justin Jenson PhD ’18 are co-first authors on the study, which appears in the latest issue of Structure.
Same problem, different approach
Jenson, for his part, has tackled the challenge of designing peptides that bind to Bcl-2 proteins using three distinct approaches. The dTERMen-based method, he says, is by far the most efficient and general one he’s tried yet.
Standard approaches for discovering peptide inhibitors often involve modeling entire molecules down to the physics and chemistry behind individual atoms and their forces. Other methods require time-consuming screens for the best binding candidates. In both cases, the process is arduous and the success rate is low.
dTERMen, by contrast, necessitates neither physics nor experimental screening, and leverages common units of known protein structures, like alpha helices and beta strands — called tertiary structural motifs or “TERMs” — which are compiled in collections like the Protein Data Bank. dTERMen extracts these structural elements from the data bank and uses them to calculate which amino acid sequences can adopt a structure capable of binding to and interrupting specific protein-protein interactions. It takes a single day to build the model, and mere seconds to evaluate a thousand sequences or design a new peptide.
“dTERMen allows us to find sequences that are likely to have the binding properties we’re looking for, in a robust, efficient, and general manner with a high rate of success,” Jenson says. “Past approaches have taken years. But using dTERMen, we went from structures to validated designs in a matter of weeks.”
Of the 17 peptides they built using the designed sequences, 15 bound with native-like affinity, disrupting Bcl-2 protein-protein interactions that are notoriously difficult to target. In some cases, their designs were surprisingly selective and bound to a single Bcl-2 family member over the others. The designed sequences deviated from known sequences found in nature, which greatly increases the number of possible peptides.
“This method permits a certain level of flexibility,” Frappier says. “dTERMen is more robust to structural change, which allows us to explore new types of structures and diversify our portfolio of potential binding candidates.”
Probing the sequence universe
Given the therapeutic benefits of inhibiting Bcl-2 function and slowing tumor growth, the Keating lab has already begun extending their design calculations to other members of the Bcl-2 family. They intend to eventually develop new proteins that adopt structures that have never been seen before.
“We have now seen enough examples of various local protein structures that computational models of sequence-structure relationships can be inferred directly from structural data, rather than having to be rediscovered each time from atomistic interaction principles,” says Grigoryan, dTERMen’s creator. “It’s immensely exciting that such structure-based inference works and is accurate enough to enable robust protein design. It provides a fundamentally different tool to help tackle the key problems of structural biology — from protein design to structure prediction.”
Frappier hopes one day to be able to screen the entire human proteome computationally, using methods like dTERMen to generate candidate binding peptides. Jenson suggests that using dTERMen in combination with more traditional approaches to sequence redesign could amplify an already powerful tool, empowering researchers to produce these targeted peptides. Ideally, he says, one day developing peptides that bind and inhibit your favorite protein could be as easy as running a computer program, or as routine as designing a DNA primer.
According to Keating, although that time is still in the future, “our study is the first step towards demonstrating this capacity on a problem of modest scope.”
This research was funded the National Institute of General Medical Sciences, National Science Foundation, Koch Institute for Integrative Cancer Research, Natural Sciences and Engineering Research Council of Canada, and Fonds de Recherche du Québec.
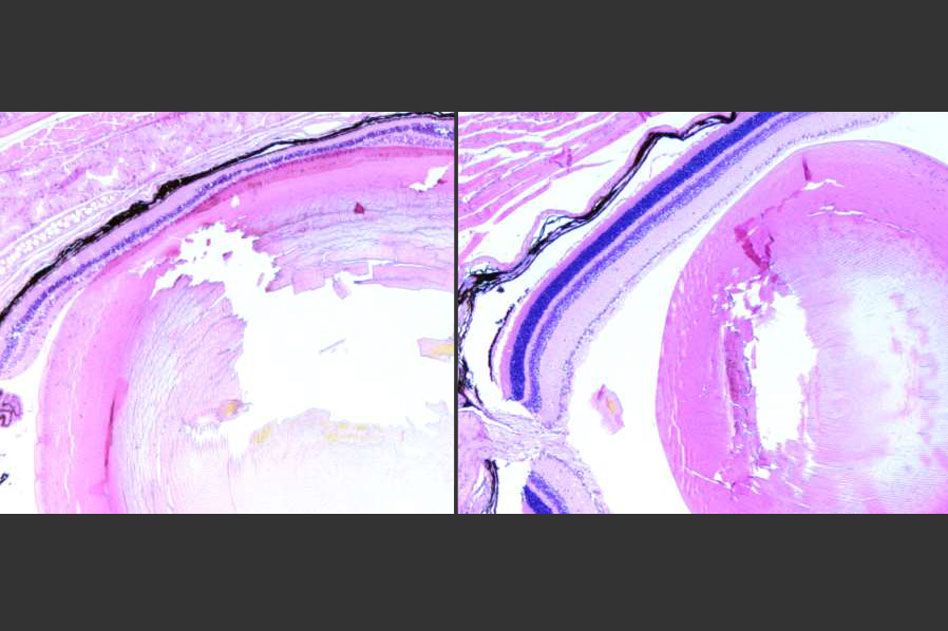
Overactive repair system promotes cell death following DNA damage by certain toxins, study shows.
Anne Trafton | MIT News Office
February 14, 2019
DNA-repair enzymes help cells survive damage to their genomes, which arises as a normal byproduct of cell activity and can also be caused by environmental toxins. However, in certain situations, DNA repair can become harmful to cells, provoking an inflammatory response that produces severe tissue damage.
MIT Professor Leona Samson has now determined that inflammation is a key component of the way this damage occurs in photoreceptor cells in the retinas of mice. About 10 years ago, she and her colleagues discovered that overactive initiation of DNA-repair systems can lead to retinal damage and blindness in mice. The key enzyme in this process, known as Aag glycosylase, can also cause harm in other tissues when it becomes hyperactive.
“It’s another case where despite the fact that inflammation is there to protect you, in some circumstances it can actually be harmful, when it’s overactive,” says Samson, a professor emerita of biology and biological engineering and the senior author of the study.
Aag glycosylase helps to repair DNA damage caused by a class of drugs known as alkylating agents, which are commonly used as chemotherapy drugs and are also found in pollutants such as tobacco smoke and fuel exhaust. Retinal damage from these drugs has not been seen in human patients, but alkylating agents may produce similar damage in other human tissues, Samson says. The new study, which reveals how Aag overactivity leads to cell death, suggest possible targets for drugs that could prevent such damage.
Mariacarmela Allocca, a former MIT postdoc, is the lead author of the study, which appears in the Feb. 12 issue of Science Signaling. MIT technical assistant Joshua Corrigan, former postdoc Aprotim Mazumder, and former technical assistant Kimberly Fake are also authors of the paper.
A vicious cycle
In a 2009 study, Samson and her colleagues found that a relatively low level of exposure to an alkylating agent led to very high rates of retinal damage in mice. Alkylating agents produce specific types of DNA damage, and Aag glycosylase normally initiates repair of such damage. However, in certain types of cells that have higher levels of Aag, such as mouse photoreceptors, the enzyme’s overactivity sets off a chain of events that eventually leads to cell death.
In the new study, the researchers wanted to find exactly out how this happens. They knew that Aag was overactive in the affected cells, but they didn’t know exactly how it was leading to cell death or what type of cell death was occurring. The researchers initially suspected it was apoptosis, a type of programmed cell death in which a dying cell is gradually broken down and absorbed by other cells.
However, they soon found evidence that another type of cell death called necrosis accounts for most of the damage. When Aag begins trying to repair the DNA damage caused by the alkylating agent, it cuts out so many damaged DNA bases that it hyperactivates an enzyme called PARP, which induces necrosis. During this type of cell death, cells break apart and spill out their contents, which alerts the immune system that something is wrong.
One of the proteins secreted by the dying cells, known as HMGB1, stimulates production of chemicals that attract immune cells called macrophages, which specifically penetrate the photoreceptor layer of the retina. These macrophages produce highly reactive oxygen species — molecules that create more damage and make the environment even more inflammatory. This in turn causes more DNA damage, which is recognized by Aag.
“That makes the situation worse, because the Aag glycosylase will act on the lesions produced from the inflammation, so you get a vicious cycle, and the DNA repair drives more and more degeneration and necrosis in the photoreceptor layer,” Samson says.
None of this happens in mice that lack Aag or PARP, and it does not occur in other cells of the eye or in most other body tissues.
“It amazes me how segmented this is. The other cells in the retina are not affected at all, and they must experience the same amount of DNA damage. So, one possibility is maybe they don’t express Aag, while the photoreceptor cells do,” Samson says.
“These molecular studies are exciting, as they have helped define the underlying pathophysiology associated with retinal damage,” says Ben Van Houten, a professor of pharmacology and chemical biology at the University of Pittsburgh, who was not involved in the study. “DNA repair is essential for the faithful inheritance of a cell’s genetic material. However, the very action of some DNA repair enzymes can result in the production of toxic intermediates that exacerbate exposures to genotoxic agents.”
Varying effects
The researchers also found that retinal inflammation and necrosis were more severe in male mice than in female mice. They suspect that estrogen, which can interfere with PARP activity, may help to suppress the pathway that leads to inflammation and cell death.
Samson’s lab has previously found that Aag activity can also exacerbate damage to the brain during a stroke, in mice. The same study revealed that Aag activity also worsens inflammation and tissue damage in the liver and kidney following oxygen deprivation. Aag-driven cell death has also been seen in the mouse cerebellum and some pancreatic and bone marrow cells.
The effects of Aag overactivity have been little studied in humans, but there is evidence that healthy individuals have widely varying levels of the enzyme, suggesting that it could have different effects in different people.
“Presumably there are some cell types in the human body that would respond the same way as the mouse photoreceptors,” Samson says. “They may just not be the same set of cells.”
The research was funded by the National Institutes of Health.
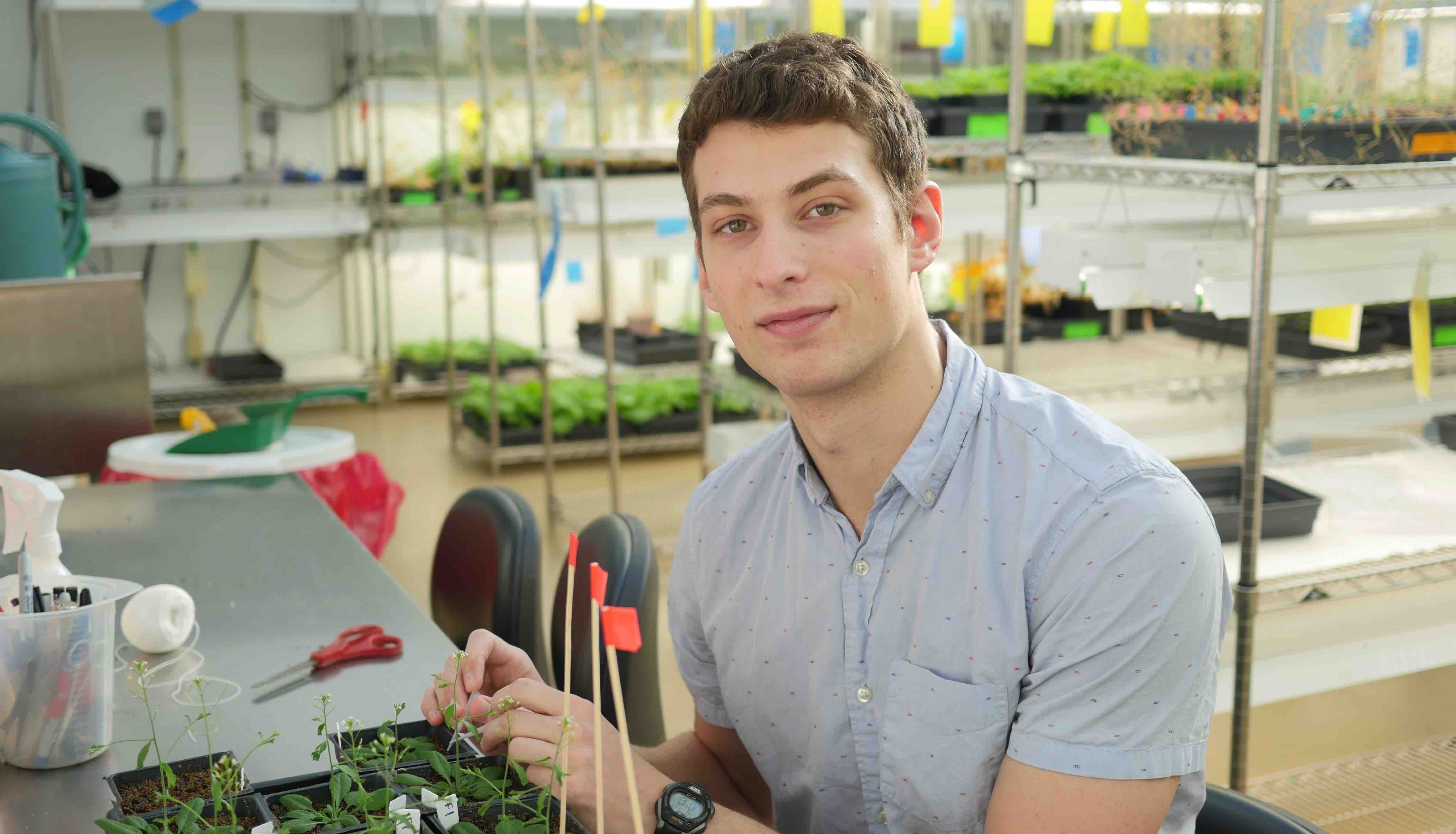

Graduate student Joe Jacobowitz analyzes new enzymes that could reveal key insights into plant reproduction.
Raleigh McElvery
January 24, 2019
Every morning, fifth-year graduate student Joe Jacobowitz takes the elevator to the seventh floor of the Whitehead Institute, passes the soil bins, “false winter” fridges, and toasty growing chambers, and enters his favorite workspace: the greenhouse. There, among the myriad of tall, stout, grass-like, and blooming plants, he attends to his organism of choice, Arabidopsis thaliana. With four simple, white petals interrupted by protruding, yellow stamens, “it looks like something that would grow in the cracks of a sidewalk,” Jacobowitz says. While you or I might pass by it and not think twice, Jacobowitz and the Weng lab hold that Arabidopsis could reveal key insights into pollen development, in particular which enzymes are critical for plant reproduction.
Jacobowitz became fascinated by enzymes as a biochemistry major at Brandeis University, studying the evolution of a single enzyme found in the deadliest form of malaria. After arriving at MIT Biology for graduate school and joining Jing-Ke Weng’s team at the Whitehead, Jacobowitz shifted his focus from biochemistry and biophysics to plant development. His work investigating the pollen-bearing chamber known as the anther represents just one facet of the Weng lab — which probes the origin and evolution of plant metabolism, as well as the small molecules plants produce to interact with their environments.
Above his lab desk, next to hand-drawn sketches and photos of friends, Jacobowitz has taped intricate microscopy images detailing the many complex stages of anther development. The pollen grains inside this structure contain the plant’s male gametes, which are transferred via wind and passersby to the female part, the pistil, of another flower. In the case of Arabidopsis, a single flower can self-pollinate and reproduce on its own, generating seeds and engendering the next generation. As the pollen grains mature, they become coated in a tough outer layer made of the material sporopollenin. This polymer, Jacobowitz explains, has helped sculpt the terrestrial ecosystem we know today.
Nearly 500 million years ago, the first plants migrated from sea to land, and eventually developed this durable coating to protect their delicate pollen grains from the stresses of living above water, such as UV radiation and desiccation. Today, researchers understand the basic sequence of events required for pollen development, but it’s been historically difficult to identify the genes involved — or even break down the resilient sporopollenin to determine its composition. In December of 2018, Weng lab postdoc Fu-Shuang Li and his team became the first to report the successful degradation of this virtually indestructible material and determine its chemical structure.
“Now that we have a better grasp of what this pollen coating looks like at a molecular level,” says co-author Jacobowitz, “we can improve our understanding of the genes that are already known to produce the pollen wall, and make predictions about new enzymes that also likely contribute.”
Jacobowitz aims to pinpoint which enzymes add certain chemical groups to sporopollenin, as well as the molecular players required for anther development. As he puts it, the general premise of his current project is to “examine genes that no one has looked at before.”
Jacobowitz spent almost a year sifting through online databases to compile a list of enzymes that could potentially play a critical role in anther development. He ordered knockout lines that eliminated each enzyme one at a time, and watched as the plants matured.
At first, nothing happened. Jacobowitz was simply rearing a bunch of normal plants. But then it occurred to him that perhaps nature had built in some redundancy, allowing plants to survive these genetic errors. If one enzyme was incapacitated, another might compensate for the loss and assume its function so development could proceed as usual.
“Even though my screens were pretty unsuccessful at first, I still enjoyed the entire process,” he recalls. “That’s when I started to realize that I really like genetics. There’s always this possibility that you’ll stumble upon a new gene, or a new function of a known gene, that no one ever suspected. That was the opposite of my undergraduate experience in biochemistry, where we drilled down into the intimate details of a single, well-studied enzyme.”With this in mind, Jacobowitz crossed two knockouts together and created a double mutant, simultaneously erasing what he suspected were two relatively similar enzymes. This time, he saw an effect — the walls of the anther began to swell, invading the space containing the pollen and preventing the grains from developing properly. He’d made a sterile plant, indicating that these two enzymes (encoded by the PRX9 and PRX40 genes, respectively) were critical for pollen development
Post-MIT, Jacobowitz is considering pursing a postdoc in genetics. He’s open to studying any organism, so plants aren’t off the table just yet.
“As humans, we rely heavily on plant-based medicines and agricultural products,” he says. “In today’s changing climate, it’s especially important recognize our dependence on plants, and put necessary resources into understanding the basic principles governing their reproductive cycle.” In fact, our own lives could depend on it.
Posted 1.24.19
Researchers refine a method to quantify protein production, show that the precision with which bacteria and eukaryotes tune their gene expression is remarkably similar.
Raleigh McElvery
December 20, 2018
January 2, 2019
Greta Friar | Whitehead Institute
December 17, 2018
CAMBRIDGE, Mass. — For hundreds of millions of years, plants thrived in the Earth’s oceans, safe from harsh conditions found on land, such as drought and UV radiation. Then, roughly 450 million years ago, plants found a way to make the move to land: They evolved spores—small reproductive cells—and eventually pollen grains with tough, protective outer walls that could withstand the harsh conditions in the terrestrial environment until they could germinate and grow into a plant or fertilize an ovule. A key component of the walls is sporopollenin, a durable polymer — a large molecule made up of many small subunits — that is absent in algae but remains ubiquitous in all land plants to this day.
Understanding the molecular composition of polymers found in nature is a fundamental pursuit of biology, with a long history tracing back to the early days of elucidating DNA and protein structures. However, the very toughness that makes sporopollenin so important for all land plants also makes it tough for researchers to study. It is extremely inert, resistant to reacting with other chemicals, including the ones researchers typically use to determine the structures of other plant biopolymers, such as polysaccharides, lignin, and natural rubber. Consequently, scientists have struggled for decades to figure out exactly what the sporopollenin polymer is made of. Now, in an article published in the journal Nature Plants on December 17, Whitehead Institute Member Jing-Ke Weng and first author and Weng lab postdoc Fu-Shuang Li, together with collaborators Professor Mei Hong and graduate student Pyae Phyo from the Massachusetts Institute of Technology (MIT) Department of Chemistry, have used innovative chemical degradation methods and state-of-the-art nuclear magnetic resonance (NMR) spectroscopy to determine the chemical structure of sporopollenin.
“Plants could not have colonized the land if they had not developed a way to withstand harsh environments,” says Weng, who is also an assistant professor of biology at MIT. “Sporopollenin helped make the terrestrial ecosystem as we know it possible.”
In addition to solving a longstanding puzzle in plant chemistry, identifying the structure of sporopollenin opens the door for its potential use in a host of other applications. Sporopollenin’s inertness is a desirable attribute to replicate in the development of, for example, medical implants such as stents, which prop open clogged arteries, to prevent negative interactions between the device and the body. It could also be a good model for durable paints and coatings, such as those used on boats, where its inertness would prevent reactions with compounds in the water and so protect the ship’s hull from environmental degradation.
Finding the shape and composition of sporopollenin was not a simple task. The first challenge was getting enough of the material to study, as pollen amounts that can be collected from most plants are minute. However, pollen from the pitch pine, Pinus rigida, is sold in bulk in China as a topping for rice cakes, so Weng used an unconventional sample collection method: He asked his parents in China to ship him copious quantities of pitch pine pollen.
A common approach to determine a complex plant polymer’s structure is to dissolve it in solutions with specific chemical compounds that will break it apart into smaller and smaller pieces from which the complete structure can be deduced. But since sporopollenin is inert and does not react with the researchers’ usual cadre of chemicals, figuring out how to break down the molecule was a key challenge.
In order to crack this problem — and make the sporopollenin dissolve more easily — Li used a specially designed grinder known as the high-energy ball mill to physically shear the tiny pollen coat into even finer pieces. Then he began testing different chemical mixtures to find ones that could break apart the sporopollenin polymer into more accessible fragments. The big breakthrough came when he tried a chemical degradation process called thioacidolysis, an acid catalyzed reaction with a pinch of a special sulfur-containing compound. This allowed Li to consistently break down 50% of the total sporopollenin polymer into small pieces, with the structure of each of these pieces resolved one by one.
To help complete the puzzle, the researchers collaborated with Mei Hong’s group in MIT’s Department of Chemistry and used magic-angle-spinning solid-state NMR spectroscopy, which can determine the chemical structures of insoluble compounds by having them interact with magnetic fields. This investigation narrowed the possible structures for sporopollenin. Combined with more chemical degradation tests to verify certain possibilities and eliminate others, it ultimately led to the complete structure.
With the structure of sporopollenin in hand, the researchers were then able to identify aspects of this unique polymer that make it such a good protective wall for spores and pollen.
A key finding was that sporopollenin molecules contain two types of cross-linkages, esters and acetals, that act like chemical clips, binding the chains of the molecule together. Other known plant polymers have only one main type of cross-link, and this unique characteristic likely provides the extreme chemical inertness of sporopollenin. Ester bonds are resistant to mildly acidic conditions, while acetals are resistant to basic conditions, meaning the molecule won’t break down in either type of environment in the wild or in the lab.
Other components of sporopollenin that the researchers found include multiple molecules known to provide UV protection, as well as fatty acids, which are water resistant and may protect spores and pollen from drought or other changes in water availability.
The researchers are now looking for differences in sporopollenin between species. Pine is not a flowering plant, but the majority of plants of interest to agriculture and medicine are, so Weng and Li are investigating how sporopollenin may have changed with the evolution of the flowering plants.
“Since I was a student, inspired by the magnificent discovery of the structure of DNA, I have been driven to discover the fundamental forms of things in nature,” Weng says. “It has been so rewarding to illuminate the structure of this crucial biopolymer in plants.”
This work was supported by the Pew Scholar Program in the Biomedical Sciences and the Searle Scholars Program, and the U.S. Department of Energy (# DE-SC0001090).
***
Jing-Ke Weng’s primary affiliation is with Whitehead Institute for Biomedical Research, where his laboratory is located and all his research is conducted. He is also an assistant professor of biology at Massachusetts Institute of Technology.
***
Full citation:
“The molecular structure of plant sporopollenin”
Nature Plants, December 17, 2018, DOI: 10.1038/s41477-018-0330-7
Fu-Shuang Li (1), Pyae Phyo (2), Joseph Jacobowitz (1,3), Mei Hong (2), Jing-Ke Weng (1,3)
1. Whitehead Institute for Biomedical Research, Cambridge, Massachusetts 02142, United States.
2. Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, United States.
3. Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, United States.
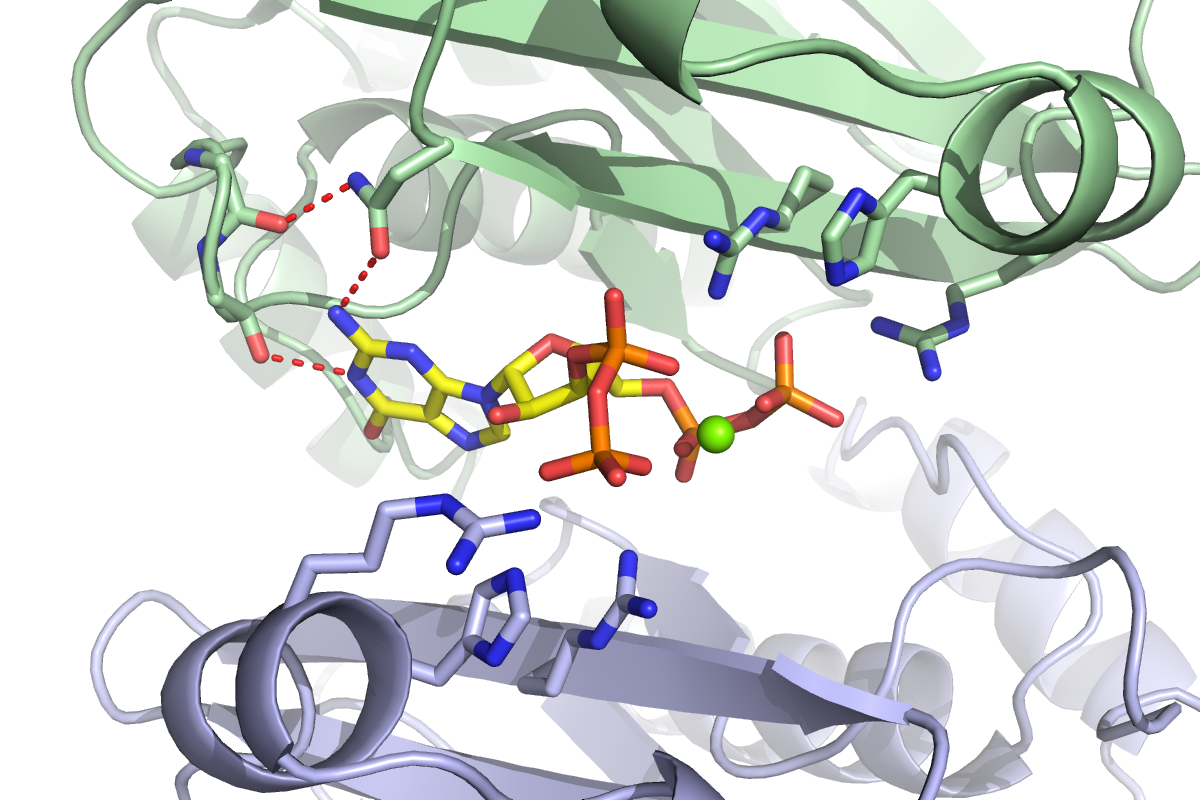
Researchers develop a method to investigate how bacteria respond to starvation and to identify which proteins bind to what they call the “magic spot” — ppGpp.
Raleigh McElvery | Department of Biology
December 17, 2018
In 1969, scientist Michael Cashel was analyzing the compounds produced by starved bacteria when he noticed two spots appearing on his chromatogram as if by magic. Today, we know one of these “magic spots,” as researchers call them, as guanosine tetraphosphate, or ppGpp for short. We also understand that it is a signaling molecule present in virtually all bacteria, helping tune cell growth and size based on nutrient availability.
And yet, despite decades of study, precisely how ppGpp regulates bacterial growth has remained rather mysterious. Delving further requires a more comprehensive list of the molecules that ppGpp binds to exert its effects.
Now, collaborators from MIT’s departments of Biology and Chemistry have developed a method to do just that, and used their new approach to pinpoint over 50 ppGpp targets in Escherichia coli — roughly half which had not been identified previously. Many of these targets are enzymes required to produce nucleotides, the building blocks of DNA and RNA. During times when the bacteria do not have enough nutrients to grow and divide normally, the researchers propose that ppGpp prevents these enzymes from creating new nucleotides from scratch, helping cells enter a dormant state.
“With small molecules or metabolites like ppGpp, it’s been difficult historically to determine which proteins they bind,” says Michael Laub, a professor of biology, a Howard Hughes Medical Institute investigator, and the senior author of the study. “This has been an intractable problem that’s held the field back for some time, but our new approach allows you to nail down the likely targets in a matter of weeks.”
Postdoc Boyuan Wang is the first author of the study, which appeares in Nature Chemical Biology on Dec. 17.
Since ppGpp was discovered nearly 50 years ago, it has been shown to suppress DNA replication, transcription, translation, and various metabolic pathways. It puts the brakes on cell growth and allows bacteria to persist in the face of starvation, stress, and antibiotics. Its influence over numerous regulatory processes has remained somewhat of a mystery, however — after all, it doesn’t just modulate a single pathway but coordinates multiple operations simultaneously to orchestrate a mass shutdown of the cell.
In order to discern which proteins ppGpp binds to effect such widespread change, the researchers built what they call “capture compounds” that contain ppGpp, allowing them to fish out its targets from bacterial extracts. These compounds included a photoreactive crosslinker that latched tightly onto the proteins of interest in the presence of light, and a biotin handle that helped the scientists pull out the proteins to identify them. Most importantly, they were joined to ppGpp in such a way that they wouldn’t interfere with its ability to bind to its targets. This method is more efficient and accurate compared to more traditional means of distinguishing ppGpp targets, which are far more arduous and lack sensitivity.
“Our approach solves these problems because you’re no longer required to do such labor-intensive protocols in order to identify ppGpp targets — and it works even in bacteria beyond E. coli,” says Wang. “Although ppGpp is common among many bacterial species, it seems to exert its effects through different mechanisms, which complicates things. Our capture compounds provide a way to unravel this diversity, and in short order.”
Although the 56 ppGpp targets Wang identified in his screen control a myriad of cellular processes, he homed in on the enzyme PurF — which initiates the biosynthesis of purine nucleotides bearing adenine and guanine bases, also known as A and G.
When bacteria are stressed or starved, they enter a dormant state to survive. But simply curbing translation and transcription is not enough; nucleotides are still being generated and will build up if their synthesis is not put on pause. Cells can build nucleotides in one of two ways: either by salvaging existing materials or starting completely from scratch. PurF kicks off the first step in the latter process leading to the A and G nucleotides. However, when ppGpp binds to PurF, it causes the enzyme to change its shape, which prevents it from doing its job, thus reducing nucleotide production in the cell.
“This is the first time that an enzyme involved in that specific pathway or function has been identified as a ppGpp target,” Wang says. “If you limit the consumption of nucleotides but not their production, the nucleotide pool is going to explode, which isn’t good for the cell. So we’ve shown that ppGpp actually addresses this problem as well.”
In addition to PurF and other enzymes required for nucleotide production, the researchers noticed that ppGpp also binds to many GTPase enzymes involved in translation. This could indicate a failsafe mechanism slowing down translation by striking multiple, similar enzymes in an almost redundant manner in the face of starvation.
As Wang continues to refine his method, he aims to increase its specificity and ensure his capture compounds bind to the exact same proteins they would inside a live cell. He also hopes to screen for ppGpp binding proteins in other bacteria, including pathogens that rely on ppGpp to survive within their hosts and propagate conditions like tuberculosis.
“This is an exciting chemical approach to better understand the function of a long-studied conserved signaling molecule in bacteria,” says Jue Wang, professor of bacteriology at the University of Wisconsin at Madison, who was not involved with the study. “Their findings and techniques are highly relevant to many other bacteria, and will greatly improve knowledge of how bacteria use this critical signaling molecule to mediate everything from surviving in the human gut to causing disease.”
Adds Laub: “We are still discovering new nucleotide-based signaling molecules in bacteria even today, and every single one of them could eventually be derivatized in a similar way to identify their binding partners.”
This research was supported by a fellowship from the Jane Coffin Childs Memorial Fund for Medical Research and a grant from the National Institutes of Health.
Snippets of RNA that accumulate in brain cells could interfere with normal function.
Anne Trafton | MIT News Office
November 27, 2018
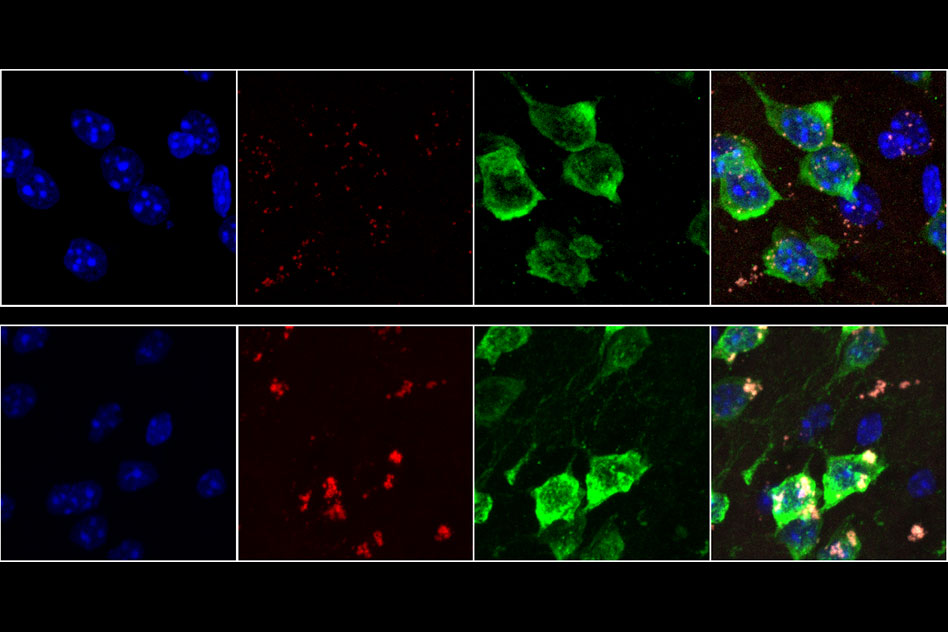
As we age, neurons in our brains can become damaged by free radicals. MIT biologists have now discovered that this type of damage, known as oxidative stress, produces an unusual pileup of short snippets of RNA in some neurons.
This RNA buildup, which the researchers believe may be a marker of neurodegenerative diseases, can reduce protein production. The researchers observed this phenomenon in both mouse and human brains, especially in a part of the brain called the striatum — a site involved in diseases such as Parkinson’s and Huntington’s.
“The brain is very metabolically active, and over time, that causes oxidative damage, but it affects some neurons more than others,” says Christopher Burge, an MIT professor of biology. “This phenomenon appears to be a previously unrecognized consequence of oxidative stress, which impacts hundreds of genes and may influence translation and RNA regulation globally.”
Burge and Myriam Heiman, the Latham Family Career Development Associate Professor of Brain and Cognitive Sciences, are the senior authors of the paper, which appears in the Nov. 27 issue of Cell Reports. Peter Sudmant, a former MIT postdoc, is the lead author of the paper, and postdoc Hyeseung Lee and former postdoc Daniel Dominguez are also authors.
A mysterious finding
For this study, the researchers used a technique developed by Heiman that allows them to isolate and sequence messenger RNA from specific types of cells. Messenger RNA carries protein-building instructions to cell organelles called ribosomes, which read the mRNA and translate the instructions into proteins by stringing together amino acids in the correct sequence.
Heiman’s technique involves tagging ribosomes from a specific type of cells with green fluorescent protein, so that when a tissue sample is analyzed, researchers can use the fluorescent tag to isolate and sequence RNA from only those cells. This allows them to determine which proteins are being produced by different types of cells.
“This is particularly useful in the nervous system where you’ve got different types of neurons and glia closely intertwined together, if you want to isolate the mRNAs from one particular cell type,” Burge says.
In separate groups of mice, the researchers tagged ribosomes from either D1 or D2 spiny projection neurons, which make up 95 percent of the neurons found in the striatum. They labeled these cells in younger mice (6 weeks old) and 2-year-old mice, which are roughly equivalent to humans in their 70s or 80s.
The researchers had planned to look for gene expression differences between those two cell types, and to explore how they were affected by age. “These two types of neurons are implicated in several neurodegenerative diseases that are aging-related, so it is important to understand how normal aging changes their cellular and molecular properties,” says Heiman, who is a member of MIT’s Picower Institute for Learning and Memory and the Broad Institute of MIT and Harvard.
To the researchers’ surprise, a mysterious result emerged — in D1 neurons from aged mice (but not neurons from young mice or D2 neurons from aged mice), they found hundreds of genes that expressed only a short fragment of the original mRNA sequence. These snippets, known as 3’ untranslated regions (UTRs), were stuck to ribosomes, preventing the ribosomes from assembling normal proteins. “While these RNAs have been observed before, the magnitude and age-associated cell-type specificity was really unprecedented,” says Sudmant.
The 3’ UTR snippets appeared to originate from about 400 genes with a wide variety of functions. Meanwhile, many other genes were totally unaffected.
“There are some genes that are completely normal, even in aged D1 neurons. There’s a gene-specific aspect to this phenomenon that is quite interesting and mysterious,” Burge says.
The findings led the researchers to explore a possible role for oxidative stress in this 3’ UTR accumulation. Neurons burn a great deal of energy, which can produce free radicals as byproducts. Unlike many other cell types, neurons do not get replaced, so they are believed to be susceptible to accumulated damage from these radicals over time.
The MIT team found that the activation of oxidative stress response pathways was higher in D1 neurons compared to D2 neurons, suggesting that they are indeed undergoing more oxidative damage. The researchers propose a model for the production of isolated 3′ UTRs involving an enzyme called ABCE1, which normally separates ribosomes from mRNA after translation is finished. This enzyme contains iron-sulfur clusters that can be damaged by free radicals, making it less effective at removing ribosomes, which then get stuck on the mRNA. This leads to cleavage of the RNA by a mechanism that operates upstream of stalled ribosomes.
“Sending neural signals takes a lot of energy,” Burge says. “Over time, that causes oxidative damage, and in our model one of the proteins that eventually gets damaged is ABCE1, and that triggers the production of 3’ UTRs.”
RNA buildup
The researchers also found the same accumulation in most parts of the human brain, including the frontal cortex, which is very metabolically active. They did not see it in most other types of human tissue, with the exception of liver tissue, which is exposed to high levels of potentially toxic molecules.
In human brain tissue, the researchers found that the amount of 3’ UTRs gradually increased with age, which fits their proposed model of gradual damage by oxidative stress. The researchers’ findings and model suggest that the production of these 3′ UTRs involves the destruction of normal mRNAs, reducing the amount of protein produced from the affected genes. This buildup of 3′ UTRs with ribosomes stuck to them can also block ribosomes from producing other proteins.
It remains to be seen exactly what effect this would have on those neurons, Burge says, but it is possible that this kind of cellular damage could combine with genetic and environmental factors to produce a general decline in cognitive ability or even neurodegenerative conditions such as Parkinson’s disease. In future studies, the researchers hope to further explore the causes and consequences of the accumulation of 3’ UTRs.
The research was funded by the National Institutes of Health and the JPB Foundation.
A new study linking RNA processing to DNA repair may open new avenues to cancer therapy.
Bendta Schroeder | Koch Institute
December 3, 2018
Mutations in the BRCA1 and BRCA2 genes pose a serious risk for breast and ovarian cancer because they endanger the genomic stability of a cell by interfering with homologous recombination repair (HR), a key mechanism for accurately repairing harmful double-stranded breaks in DNA. Without the ability to use HR to fix double-stranded breaks, the cell is forced to resort to more error-prone — and thus more cancer-prone — forms of DNA repair.
The BRCA1 and BRCA2 genes are not the only genes whose mutations foster tumorigenesis by causing an inability to repair DNA double strand breaks by HR. Mutations in twenty-two genes are known to disrupt HR, giving rise to tumors with what researchers call “BRCAness” characteristics. All but one of these BRCAness genes are known to be directly involved in the HR pathway.
The one exception, CDK12, is thought to facilitate a set of different processes altogether, involving how RNA transcripts are elongated, spliced and cleaved into their mature forms. While the connection between this RNA-modulating gene to DNA repair remained poorly understood, the identification of CDK12 as a BRCAness gene piqued significant clinical interest.
The researchers who pinpointed this connection, Sara Dubbury and Paul Boutz, both work in the laboratory of Phillip Sharp, Institute Professor, professor of biology, and member of the Koch Institute for Integrative Cancer Research. In a study appearing online in Nature on Nov. 28, they describe how they discovered a previously unknown mechanism by which CDK12 enables the production of full-length RNA transcripts and that this mechanism was especially critical to maintain functional expression of the other BRCAness genes.
When the researchers knocked out expression of CDK12, mouse stem cells showed many signs of accumulating DNA damage that prevented DNA replication from going forward, classic indications of a BRCAness phenotype. To identify what roles CDK12 may play in regulating gene expression, the researchers turned to RNA sequencing to determine which genes had increased or decreased their overall expression.
To their surprise, only genes activated by p53 and early differentiation (side effects of accumulating unrepaired DNA damage and BRCAness in mouse stem cells) accounted for the lion’s share of changes to RNA transcription. However, when the researchers instead focused on the types of RNAs transcribed, they found that many genes produced unusually short transcripts when CDK12 was absent.
Not every stretch of DNA in a gene makes it into the final RNA transcript. The initial RNA from a gene often includes sections, which researchers call “introns,” that are cut out of transcript, the discovery that earned Sharp the 1993 Nobel Prize in Physiology or Medicine and the remaining sections. “Exons,” are spliced together to form a mature transcript (mRNA). Alternately, an intronic polyadenylation (IPA) site may be activated to cleave away the RNA sequence that follows it preventing intron removal and generating a prematurely shortened transcript. These processes allow the same gene to produce alternate forms of messenger RNA (mRNA), and thus be translated into different protein sequences.
Surprisingly CDK12 knockout cells produced significantly more IPA-truncated transcripts genome-wide, while full-length transcripts for the same genes were reduced. These shortened mRNAs can vary greatly in their stability, their ability to be translated into protein, and their protein function. Thus, even while a gene may be actively transcribed, its translation into functional proteins can be radically altered or depleted by IPA activation.
While this observation began to illuminate CDK12’s role in regulating mRNA processing, what remained puzzling was why CDK12 loss affected the HR pathway so disproportionately. In investigating this question, Dubbury and Boutz found that BRCAness genes were overrepresented as a group among those genes that have increased IPA activity upon CDK12 loss.
Additionally, while CDK12 suppresses IPA activity genome-wide, 13 of the other 21 BRCAness genes were found to be particularly vulnerable to CDK12 loss, in part, because they possess multiple high-sensitivity IPA sites, which have a compound effect in decreasing the total amount of full-length transcripts. Moreover, because multiple CDK12-senstive BRCAness genes operate in the same HR pathway, the researchers believe that the disruption to HR repair of double-stranded DNA breaks is amplified.
CDK12 mutations are found recurrently in prostate and ovarian cancer patients, making them an attractive diagnostic and therapeutic target for cancer. However, not enough is known about CDK12 to distinguish between true loss-of-function mutations and so-called “passenger mutations” with no functional consequence.
“The ability to identify patients with true loss-of-function mutations in CDK12 would enable clinicians to label a new cohort of patients with bona fide BRCAness tumors that could benefit from certain highly effective and targeted chemotherapeutics against BRCAness, such as PARP1 inhibitors,” says Dubbury, a former David H. Koch Fellow.
Dubbury and Boutz were able to confirm that IPA sites in key BRCAness genes were also used more frequently upon CDK12 loss in human tumor cells using RNA sequencing data from prostate and ovarian tumor patients with CDK12 mutations and by treating human prostate adenocarcinoma and ovarian carcinoma cells with a CDK12 inhibitor. This result suggests that the CDK12 mechanism observed in mouse cell lines is conserved in humans and that CDK12 mutations in human ovarian and prostate tumors may promote tumorigenesis by increasing IPA activity and thus functionally attenuating HR repair.
“These results not only give us a better understanding how CDK12 contributes to BRCAness, they also may have exciting potential impact in the clinic,” Dubbury says. “Currently available diagnostic techniques could be used to probe the usage of IPA sites found in this study to rapidly screen for patients with true loss-of-function CDK12 mutations, who would respond to BRCAness-targeted treatments.”
Paul Boutz, a research scientist in the Sharp Lab, is co-first author of the study, and has plans to follow-up many of these implications for ovarian and prostate cancer his lab at the University of Rochester School of Medicine and Dentistry.
“CDK12 provides a remarkable example of how factors that control the processing of RNA molecules can function as master regulators of gene networks, and thereby profoundly affect the physiology of both normal and cancerous cells,” he says.
Phil Sharp, the senior author on the work, says “Sara’s and Paul’s surprising discovery that CDK12 suppresses intronic polyadenylation has implications for fundamental new insights into gene structure as well as for control of cancer.”
