Eva Frederick | Whitehead Institute
April 20, 2021
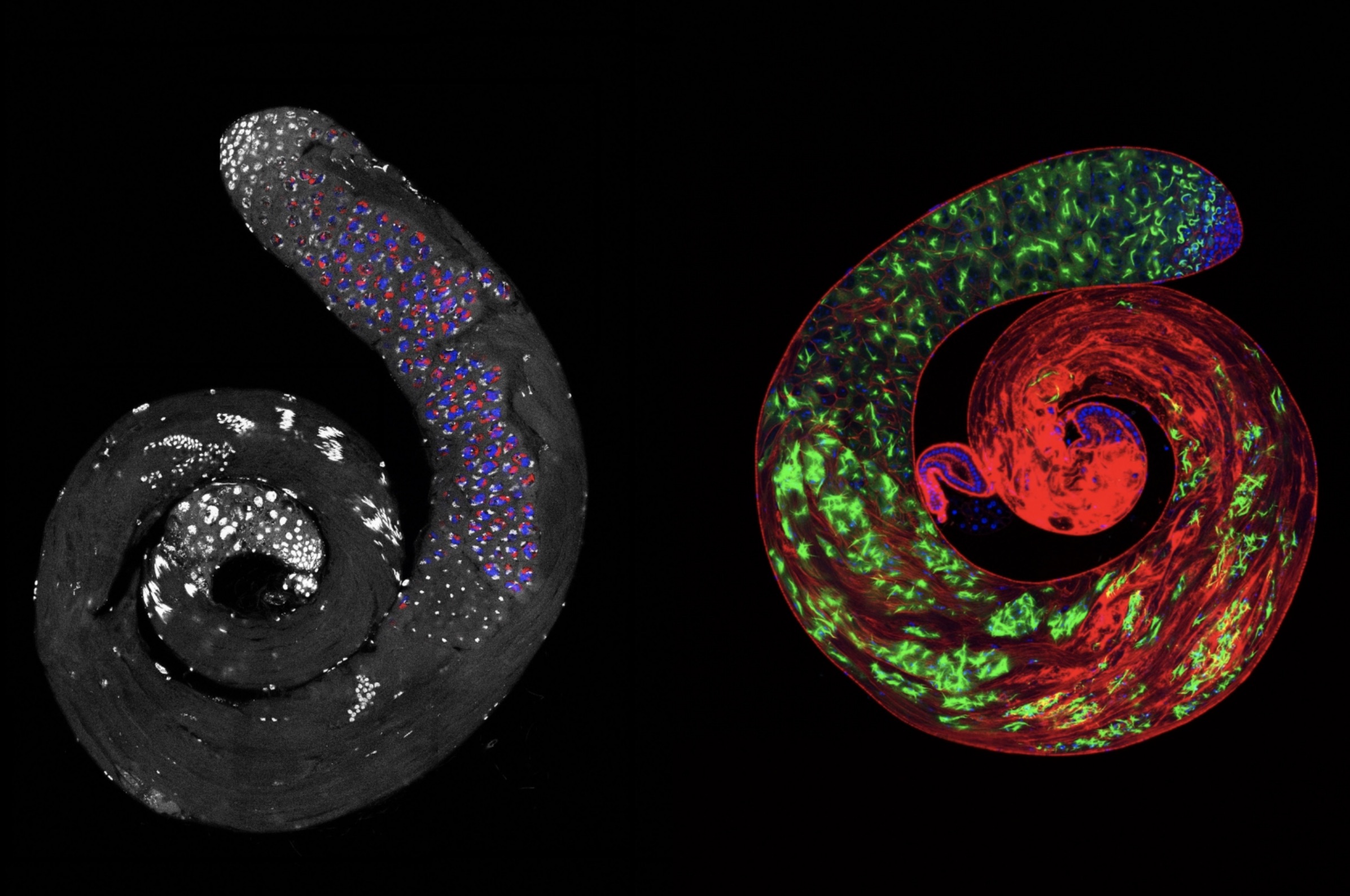
Planarians are small water-dwelling worms known for their regenerative capacity. If you chop one into ten pieces, you’ll end up with ten fully-formed worms.
While humans have pools of specialized stem cells that can create our regenerative body parts like hair and skin, these worms owe their regenerative superpowers to a special kind of stem cell called a neoblast. At least some of these cells are “pluripotent,” meaning that they can divide to create almost any cell type in a worm’s body at any time. Neoblasts are actually the only dividing cells in planarians — fully committed cells like those in the eyes or intestines cannot divide again.
“The big question for us is, how does a neoblast go from being able to make anything, to making one particular thing?” says Amelie Raz, a postdoctoral researcher at Whitehead Institute who conducted her graduate research in the lab of Whitehead Institute Member Peter Reddien. “How do they go from being able to make anything in the body to being, say, an intestine cell that’s going to stay an intestine cell until it dies?”
Now, in a paper published online April 20 in the journal Cell Stem Cell, researchers at Whitehead Institute lay out a new model for how these stem cells commit to their fates and go on to create fully differentiated cells. The process of cellular differentiation is often viewed as a hierarchy, with one special stem cell at the top which can take a number of potential paths to arrive at a specialized state. This is generally thought to take place over a series of cell divisions in which each generation’s fate is gradually restricted.
“We’re proposing something happens that is very different from the conventional view,” says senior author Reddien, who is also a professor of biology at Massachusetts Institute of Technology and an investigator with the Howard Hughes Medical Institute. “We think that stem cells can make broad jumps in state without going through a series of fate-restricting divisions. We call it the single-step fate model.”
In the new model, neoblasts that are on a path toward creating skin cells or intestine cells can produce progeny cells that can switch fates to create cells of other types. The work is a step in the long road to understanding these worms’ regenerative capacities, and could possibly inform regenerative medicine approaches far in the future.
“The ability of planarian stem cells to essentially switch their fate is really, really powerful,” says Raz, the first author of the paper. “Obviously this is a long way off, but theoretically the concept of stem cell fate switching could be applied to regenerative medicine, with human stem cell programming.”
Upturning the hierarchy
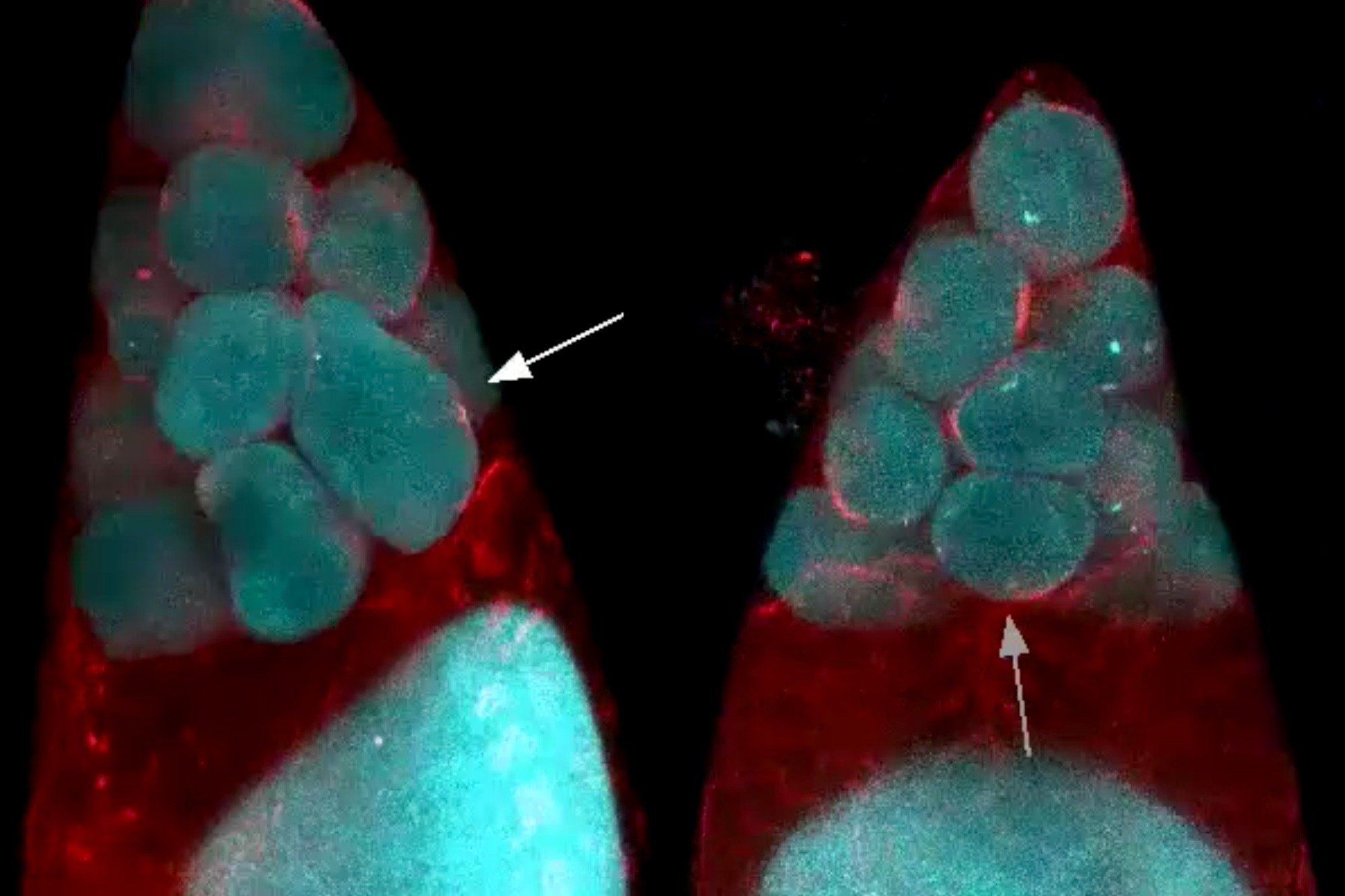
Neoblasts can be sorted into many “classes.” For example, one class of neoblasts contains all the materials to make skin cells, and others have the necessary toolkit to form the worms’ primitive kidneys or their intestines. According to the hierarchical model, these specialized neoblasts are intermediaries between a pluripotent cell at the top of the hierarchy, and the non-dividing body cells.
“You can imagine that the special cell at the top is a blank slate with no predisposition towards any cell type — it can make anything,” says Raz. “This is how we’ve often imagined development works.”
But Raz, Reddien and Omri Wurtzel, a former postdoc in the Reddien lab now at Tel Aviv University, started to question this assumption after noticing a few mysterious properties of planarian cells.
First of all, researchers have observed in the past that when a planarian is treated with radiation to kill all existing stem cells, a single neoblast can rescue the animal by forming a colony containing many different classes of neoblasts. If, as previous theories suggested, there was a single class of neoblast that gave rise to all these types, Raz and Reddien reasoned that that class should be a common resident in every colony that formed. After creating many of these colonies and analyzing their composition, however, the researchers saw that this was not the case. “For every class we looked at, there were plenty of colonies that lacked that class altogether,” says Reddien. “There was no unique class present in all colonies.”
Another sticking point: the researchers began to realize that, when applying the hierarchy model, the math of planarian cell divisions and potency just didn’t add up. In a prior cell transplantation study, the Reddien lab found that many of the neoblasts they tested were pluripotent —in this study they found that proportion to be larger than what they would expect if only non-specialized neoblasts were pluripotent. “When you add up all the different kinds of specialized neoblasts, it’s at least three quarters of the neoblast population, and almost certainly higher than that.” says Raz. Therefore, the researchers wondered if some specialized neoblasts could be pluripotent as well.
Another study from the Reddien lab showed that skin-specialized neoblasts did not retain skin fate through more than one cell division. Also, in about half of all cell divisions in planarians, the two daughter cells will be different from one another. This raised the possibility that specialized neoblasts can divide asymmetrically as a possible route to stem cells changing fate.
Furthermore, the timeline for regeneration was off — the rate at which planarians were able to regrow body parts didn’t allow for several rounds of fate-restricting divisions.
After conducting experiments to study these different situations, Raz, Wurtzel, and Reddien were able to create a case for their new model of cell differentiation. “What we think is happening is that planarians have a ton of plasticity in their general stem cell population, where individual cells can move in and out of different specialized stages through the process of cell division in order to give rise to what is required,” Raz says.
“This is just the beginning of exploring this process, even though we’ve been studying it for many years,” Reddien says. “Focusing on the model, we’re suggesting that the cells can choose one fate, and then through the process of a division with an asymmetric outcome, one of the daughter cells can now divide again and choose a different fate. That fate switching process might be fundamental to explaining pluripotency.”
Reddien’s lab will continue investigating the mechanisms of neoblast fate specification, including how specialization lines up with the timing of the cell cycle.
“Understanding the structure of cell lineage and how fate choices are made is fundamental to understanding adult stem cell biology, and how in the context of injury and repair, new cells can be brought about,” says Reddien. “Do they have to go through long, complex lineage trajectories? Or can they make big jumps in state from stem cells to the final state? How flexible is that? All of these things have potential implications for understanding stem cell biology broadly, and we hope that the work will highlight some of these mechanisms and provide opportunities to explore general principles in the future.”