Poise or Pause: researchers expand understanding of transcription factor’s role with newly discovered conformation
Lillian Eden | Department of Biology
February 23, 2024
New research from the Vos Lab in the Department of Biology at MIT reveals the dynamic nature of elongation factor protein key for regulating early stages of gene expression.
Transcription, the process of copying RNA from DNA, is a critical first step for cells to create proteins. The enzyme responsible for transcription is a motor protein called RNA polymerase.
When an RNA polymerase transcribes a gene, it will begin elongating the mRNA and will then, often, pause.
From there, the RNA polymerase will either return to elongating the mRNA or it will get stuck. For the latter occurrence, the mRNA and subsequent protein will never be made: the polymerase will go somewhere else, or restart transcription on the same gene and get stuck again.
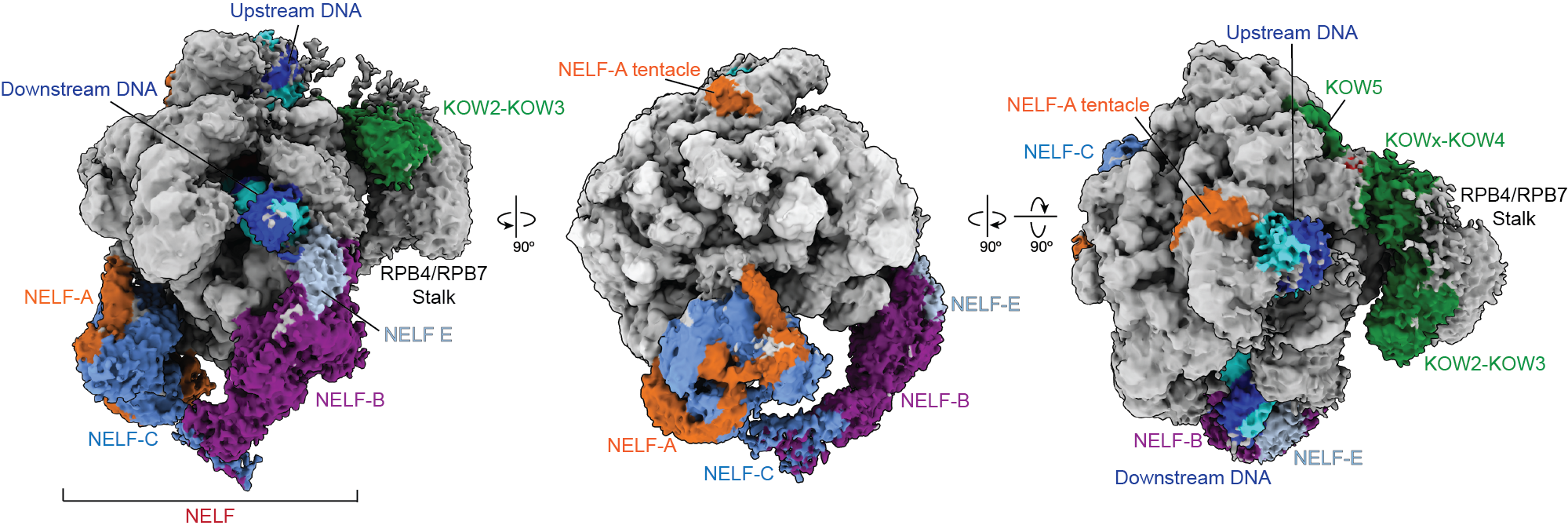
Pausing is thought to be governed by a protein called Negative Elongation Factor, NELF, and DRB-sensitivity inducing factor, DSIF. Previous research suggested that NELF stably clamps down onto RNA polymerase to stall the elongation process and prevent the polymerase from moving. That model contradicted cell-based experiments, however, which indicated that NELF is somehow still attached to polymerase after transcription resumes.
New research from the Vos Lab in the Department of Biology at MIT published today in Molecular Cell reveals that NELF isn’t merely an on-off switch for transcription. Instead, NELF can change into a distinct conformation that allows the polymerase to resume transcription. The researchers dubbed this distinct conformation NELF’s “poised” state.
RNA polymerase pausing, sometimes for minutes at a time, is thought to be an important gene expression checkpoint; more than half of genes exhibit pausing, although many questions remain about the role of pausing in gene expression. Understanding both how and why the process is occurring, down to the atomic level, and what components are involved, is key to understanding how cells function, both individually and as part of an organism.
“It’s a very central question to biological research, and we still don’t fully understand it because it’s such a complex process,” says first author Bonnie G. Su, a graduate student in the Vos lab. “The bigger picture is: how does the cell decide what resources to allocate to certain biological processes? This finding might help us answer questions like that.”
To visualize the two distinct conformations of NELF and polymerase, the researchers used a combination of biochemical and structural approaches. The previous understanding of proximal pausing was based on Cryo-Electron Microscopy images of the static complex. Cryo-EM is a powerful microscopy technique that involves freezing samples and imaging them, and that approach had captured polymerase in its paused state.
Using the core Cryo-EM facility available at MIT.nano, Su instead added the necessary components for the polymerase to transcribe, and gathered structural data on an actively transcribing complex —allowing, for the first time, a stepwise visualization of how proximal pausing occurs.
“What we found is that NELF, which we always thought of as static, can actually move around,” Su says. “This has updated our understanding of what pausing is, and how early gene regulation happens.”
The structural results also provide an explanation for how polymerase may be cycling between the two states—and how one form of NELF may be forcing polymerase to pause, while the newly discovered form allows polymerase to resume transcription.
It’s still unclear what triggers NELF to transition from the paused state to the poised state, and many questions remain about how polymerase is regulated, according to senior author Seychelle Vos, the Robert A. Swanson (1969) Career Development Professor of Life Sciences and HHMI Freeman Hrabowski Scholar. RNA polymerase can be associated with and is known to be regulated by a large repertoire of proteins.
“We’re trying to see if we can actually lock the complex in the paused state by adding additional factors,” Vos says. “We’re also pursuing whether sequence context is affecting pausing behavior—how or if the sequence of DNA may be causing polymerase to pause.”
