With a minor in literature and environmental sustainability, the biology alumna considers perspectives from Charles Darwin to Annie Dillard.
Lillian Eden | Department of Biology
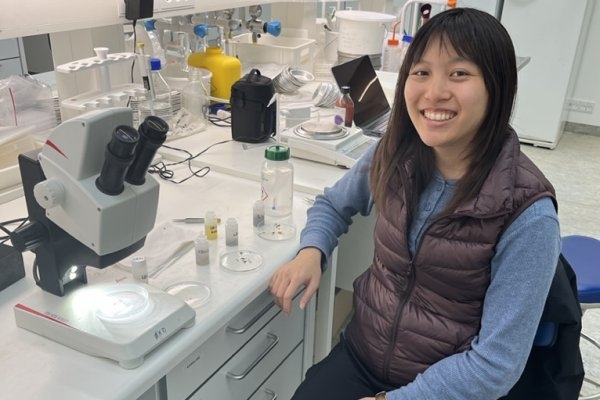
Growing up in Los Angeles about 10 minutes away from the Ballona Wetlands, Andrea Lo ’21 has long been interested in ecology. She witnessed, in real-time, the effects of urbanization and the impacts that development had on the wetlands.
“In hindsight, it really helped shape my need for a career — and a life — where I can help improve my community and the environment,” she says.
Lo, who majored in biology at MIT, says a recurring theme in her life has been the pursuit of balance, valuing both extracurricular and curricular activities. She always felt an equal pull toward STEM and the humanities, toward wet lab work and field work, and toward doing research and helping her community.
“One of the most important things I learned in 7.30[J] (Fundamentals of Ecology) was that there are always going to be trade-offs. That’s just the way of life,” she says. “The biology major at MIT is really flexible. I got a lot of room to explore what I was interested in and get a good balance overall, with humanities classes along with technical classes.”
Lo was drawn to MIT because of the focus on hands-on work — but many of the activities Lo was hoping to do, both extracurricular and curricular, were cut short because of the pandemic, including her lab-based Undergraduate Research Opportunities Program (UROP) project.
Instead, she pursued a UROP with MIT Sea Grant, working on a project in partnership with Northeastern University and the Charles River Conservancy with funding support from the MIT Community Service fund as part of STEAM Saturday.
She was involved in creating Floating Wetland kits, an educational activity directed at students in grades 4 to 6 to help students understand ecological concepts,the challenges the Charles River faces due to urbanization, and how floating wetlands improve the ecosystem.
“Our hope was to educate future generations of local students in Cambridge in order for them to understand the ecology surrounding where they live,” she says.
In recent years, many bodies of water in Massachusetts have become unusable during the warmer months due to the process of eutrophication: stormwater runoff picks up everything — from fertilizer and silt to animal excrement — and deposits it at the lowest point, which is often a body of water. This leads to an excess of nutrients in the body of water and, when combined with warm temperatures, can lead to harmful algal blooms, making the water sludgy, bright green, and dangerously toxic.
The wetland kits Lo worked with were mini ecosystems, replicating a full-sized floating wetland. One such floating wetland can be seen from the Longfellow Bridge at one end of MIT’s campus — the Charles River floating wetland is a patch of grass attached to a buoy like a boat, which is often visited by birds and inhabited by much smaller critters that cannot be seen from the shore.
The Charles River floating wetland has a variety of flora, but the kits Lo helped present use only wheat grass because it is easy to grow and has long, dangling roots that could penetrate the watery medium below. A water tray beneath the grass — the Charles river of the mini ecosystem — contains spirulina powder for replicating algae growth and daphnia, which are small, planktonic crustaceans that help keep freshwater clean and usable.
“This work was really fulfilling, but it’s also really important, because environmental sustainability relies on future generations to carry on the work that past generations have been doing,” she says. “MIT’s motto is ‘mens et manus’ — education for practical application, and applying theoretical knowledge to what we do in our daily lives. I think this project really helped reinforce that.”
Since 2021, Lo has been working in Denmark in a position she learned about through the MIT-Denmark program.
She chose Denmark because of its reputation for environmental and sustainability issues and because she didn’t know much about it except for it being one of the happiest countries in the world, often thought of synonymously with the word “hygge,” which has no direct translation but encapsulates coziness and comfort from the small joys in life.
“At MIT, we have a very strong work-hard, play-hard culture. I think we can learn a lot from the work-life balance that Denmark has a reputation for,” she says. “I really wanted to take the opportunity in between graduation and whatever came after to explore beyond my bubble. For me, it was important to step back, out of my comfort zone, step into a different environment — and just live.”
Currently, her personal project is comparing the conditions of two lagoons on the island of Fyn in Denmark. Both are naturally occurring, but in different states of environmental health.
She’s been doing a mix of field work and lab work. She collects sediment and fauna samples using a steel corer, or “butter stick” in her lab’s slang. In the same way that one can use a metal tube-shaped tool to remove the core of an apple, she punches the steel corer into the ground, removing a plug of sample. She then sifts the sample through 1 millimeter mesh, preserves the filtered sample in formalin, and takes everything back to the lab.
Once there, she looks through the sample to find macrofauna — mollusks, barnacles, and polychaetes, a bristly-looking segmented worm, for example. Collected over time, sediment characteristics like organic matter content, sediment grain size, and the size and abundance of macrofauna, can reveal trends that can help determine the health of the ecosystem.
Lo doesn’t have any concrete results yet, but her data could help researchers project the recovery of a lagoon that was rehabilitated using a technique called managed realignment, where water is allowed to reclaim areas where it was once found. She says she’s glad she gets a mix of field work and lab work, even on Denmark’s stormiest days.
“Sometimes there are really cold days where it’s windy and I wish I was in the lab, but, at the same time, it’s nice to have a balance where I can be outside and really be hands-on with my work,” she says.
Reflecting her dual interests in the technical and the innovative, she will be back in the Greater Boston area in the fall, pursuing a master of science in innovation and management and an MS in civil and environmental engineering at the Tufts Gordon Institute.
“So much has happened and changed due to the pandemic that it’s easy to dwell on what could’ve been, but I tell myself to be optimistic and take the positive aspects that have come out of the circumstances,” Lo says. “My opportunities with the Sea Grant, MISTI, and Tufts definitely wouldn’t have happened if the pandemic hadn’t happened.”