Spying on enzymes while they perform chemical reactions could help treat gut ailments
Raleigh McElvery
Humans breathe oxygen, but many microbes deep within in our gut don’t have access to this precious resource. Instead, they breathe sulfur compounds, releasing hydrogen sulfide in the process. This colorless gas is best-known for its rotten stench, but inside the human colon it has been linked to a thinner mucus barrier, and ailments such as inflammatory bowel disease, Crohn’s disease, ulcerative colitis, and colorectal cancer. In order to develop potential treatments, researchers are probing how microbes create hydrogen sulfide and which molecules they use.
To help further these efforts, Catherine Drennan’s lab and Heather Kulik’s lab at MIT collaborated with Emily Balskus’ lab at Harvard University to investigate the structure and mechanism of an enzyme that’s critical for hydrogen sulfide production: isethionate sulfite-lyase (IslA). The team examined IslA while it was bound to a metabolite that’s readily available in the gut — and revealed how the bacterium Bilophila wadsworthia uses this interaction to help generate the hydrogen sulfide precursor called sulfite. The researchers then compared IslA’s binding behavior to other enzymes in the same family, in order to better understand how these enzymes have evolved to perform challenging chemistry on a wide variety of molecules. Their findings were published on Mar. 26 in the journal Cell Chemical Biology.
“Although abundant, sulfide-producing bacteria are not well understood,” says Drennan, a professor of biology and chemistry and a Howard Hughes Medical Institute investigator. “By characterizing the enzymes in these bacteria that are responsible for sulfur metabolism, we can develop therapeutic strategies to limit production of hydrogen sulfide that can lead to disease.”
Although researchers have been studying bacterial sulfur respiration for decades, IslA was only recently identified. This enzyme breaks the bond between a carbon atom and a sulfur atom in a compound called isethionate, which is a prevalent metabolite in the human body. In doing so, IslA releases the sulfite that bacteria such as B. wadsworthia use to produce hydrogen sulfide.
IslA is a member of a large family of enzymes, known as glycyl radical enzyme (GREs). Scientists can learn a lot from examining the way GREs bind to other molecules, according to Christopher Dawson PhD ’20, the study’s co-first author.
GREs contain a binding site (or “active site”) where they latch onto their respective substrates to perform chemical reactions. “Understanding GREs better will aid in drug design efforts to combat the deleterious effects of some of these enzymes,” Dawson says. “It will also help to engineer enzymes that perform diverse, challenging reactions to expand the toolkit for chemical synthesis.”
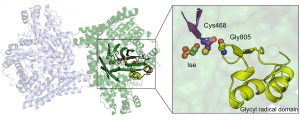
To this end, Dawson wanted to compare IslA’s active site — where it binds to isethionate to break the C-S bond — to other enzymes in the GRE family. He used X-ray crystallography to visualize this interaction at the level of individual atoms. The GREs he examined shared similar “barrel-like” structures in their active sites, but used these core features in different ways, depending on the substrates they bound. For instance, isethionate bound higher in IslA’s active site compared to the way other GREs bind their respective substrates. While this aberrant binding behavior is quite unique — even among GREs — another group had found something similar when they elucidated IslA’s structure in a different bacterium. And, the Drennan lab suspects this pattern could be prevalent in other classes of enzymes as well.
Next, Dawson and his colleagues wanted to investigate how IslA goes about cleaving the C-S bond once the enzyme has bound to isethionate. Others had predicted this process would occur via a “migration” reaction. In that scenario, the sulfite leaving group first migrates to another carbon atom and then that C-S bond is cleaved to release it. However, after co-first author Stephania Irwin generated multiple IslA variants, the Kulik lab performed computational analyses, and the researchers completed structural comparisons, the team concluded that migration was not occurring. Instead, IslA appeared to be performing an “elimination” reaction that severed the C-S bond without forming another one via migration.
Now that they know more about IslA — and GREs in general — the researchers hope their insights will aid drug design.
“Understanding how pathogens use enzymes like these to extract sulfite from their hosts and fuel hydrogen sulfide production has very clear therapeutic implications,” Dawson says. “And that’s one of the things I like best about this story.”
Citation
“Molecular Basis of C-S Bond Cleavage in the Glycyl Radical Enzyme Isethionate Sulfite-Lyase”
Cell Chemical Biology, online March 26, 2021,
DOI: 10.1016/j.chembiol.2021.03.001
Christopher D. Dawson, Stephania M. Irwin, Lindsey R. F. Backman, Chip Le, Jennifer X. Wang, Vyshnavi Vennelakanti, Zhongyue Yang, Heather J. Kulik, Catherine L. Drennan, and Emily P. Balskus