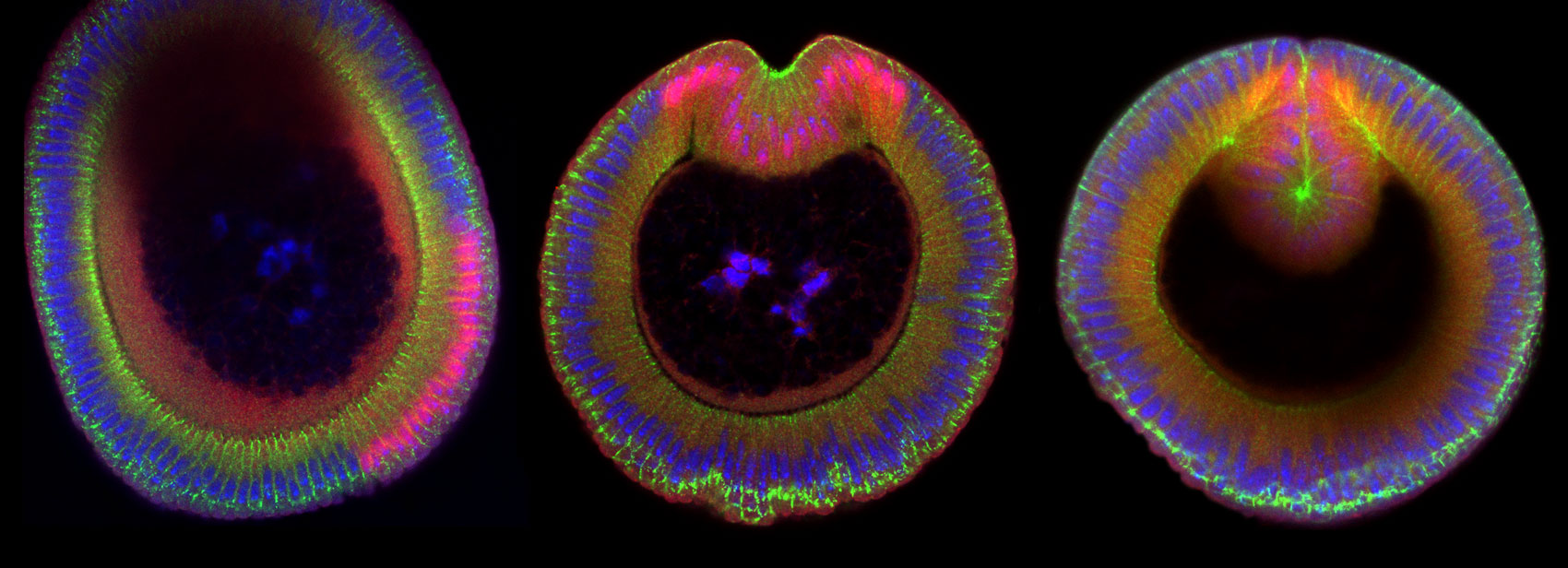
Patterns of myosin and F-actin proteins across developing embryos promote tissue folding and shape new life.
Raleigh McElvery
June 14, 2021
Virtually all multicellular organisms, including humans, begin as a single cell that rapidly divides and begets trillions of others. These cells work together, stretching, squishing, and migrating to sculpt organs and tissues. In the case of the fruit fly embryo, it only takes a few hours for life to take shape. First, the multiplying cells form an oblong sphere akin to a football. Then, mechanical forces cause a band of cells along the “belly” side of the developing fly to furrow inwards. These “mesoderm” cells form a new layer that will later give rise to muscles. Although the folding process transpires in less than 20 minutes, it’s crucial for determining where the cells will go and what roles they will assume.
Scientists previously identified two important proteins that generate the force needed to fold the tissue. The first, myosin, has a characteristic rod shape with feet hanging off both ends. It can “walk” along the cell’s inner scaffolding, composed of a second, rope-like protein called filamentous actin (F-actin). As it walks, myosin tugs on the F-actin and constricts the tissue. Researchers are probing the distribution of myosin and F-actin across the developing embryo, an important step towards understanding how these proteins drive constriction in the proper places to fold the tissue.
Myosin appears in a gradient across the belly and back of the developing fly. Since myosin and F-actin work together, many scientists assumed they would display the same pattern. However, new work from MIT’s Department of Biology and Department of Mathematics suggests otherwise. The study, published in the journal Development, shows how gene expression patterns dictate a unique distribution of F-actin across the mesoderm, which exhibits peaks and valleys. In combination with the myosin present, this F-actin pattern causes the cells to stretch, squish, or maintain their shape in just the right places to bend the tissue.
“We’ve known for decades that mechanical proteins like myosin and F-actin regulate tissue curvature during development,” says Adam Martin, an associate professor of biology and the study’s senior author. “But what hasn’t been appreciated is the extent to which these two proteins are intricately patterned during the tissue folding process. Our finding that F-actin has a different pattern than myosin was quite surprising.”
The researchers, led by graduate student Marlis Denk-Lobnig, began by focusing on two well-known transcription factor proteins, Twist and Snail, which bind to DNA to control gene expression. These transcription factors are known to dictate cell shape and fate during tissue folding, and Denk-Lobnig wondered how they affected F-actin levels.
By imaging live and fixed cells, the researchers observed that Snail and Twist drove a different pattern of F-actin density across the mesoderm compared to previously described myosin gradients. Two to three hours after the fruit fly eggs are laid, Snail depletes F-actin levels across the mesoderm. But, as Twist activates its transcriptional targets, F-actin and myosin levels rise in a subset of the mesoderm cells along the belly of the developing fly — constricting them and folding that swath of the tissue. The more F-actin and myosin a cell contains, the more compressed and wedge-shaped it becomes.
Denk-Lobnig also targeted another protein, RhoA, that tunes F-actin and myosin levels. RhoA activation is ultimately controlled by the ratio of two other molecules, C-GAP and RhoGEF2. The researchers adjusted the levels of C-GAP and RhoGEF2 in live cells, and watched the subsequent changes in myosin and F-actin distribution in real time.
To continue disentangling the effects of each protein on tissue curvature, they leveraged a computer simulation of a developing embryo designed by Associate Professor of Mathematics Jörn Dunkel, former grad student Pearson Miller PhD ’20, and postdoctoral researcher Jan Totz. The model allowed the team to adjust patterns of force and protein activity, in order to determine how the changes that they’d witnessed in real embryos affected tissue shape.
“The main takeaway is that you need this elegant coordination between cells during development,” Denk-Lobnig says. “We’ve shown how force generation patterns change the shape of individual cells — and how this leads to shape changes across entire tissues.”
Top image: Cross sections of three fruit fly embryos undergoing tissue folding. Nuclei are in blue, the transcription factor Snail is in red, and the junctional protein Armadillo is in green. Credit: Marlis Denk-Lobnig.
Video: An early stage fruit fly embryo has a band of cells on its surface that furrows inward to form a fold. Credit: Marlis Denk-Lobnig.
Citation:
“Combinatorial patterns of graded RhoA activation and uniform F-actin depletion promote tissue curvature”
Development, online June 14, DOI: 10.1242/dev.199232
Marlis Denk-Lobnig, Jan F. Totz, Natalie C. Heer, Jörn Dunkel, and Adam C. Martin
