Eva Frederick | Whitehead Institute
July 20, 2022
Multicellular organisms evolved over millennia into a dazzling array of differently adapted creatures. With each generation, tiny worms, lavishly plumed birds, and even humans must create themselves anew from a single cell. To do so, they require a plan.
“How that multifunctional body plan is created is one of the deepest questions in developmental biology,” said Zak Swartz, until recently a postdoctoral researcher in the lab of Whitehead Institute Member Iain Cheeseman. “How do you take a single cell and pattern into a body that has different functions and features along it?”
Whitehead Institute researchers are tackling this question through a variety of different lenses. Researchers in Iain Cheeseman’s lab, including Swartz, have delved into the mysterious forces that underlie the polarity of an organism’s first cell. For the lab led by Pulin Li, research comes in at a later stage of development, when multiple cells combine to form a tissue and must communicate with each other to become an organized whole. Work on regeneration in Peter Reddien’s lab shows how some creatures can access their body blueprint throughout their lives to repair nearly any injury, and Yukiko Yamashita’s group studies how organisms pass on their body blueprints to their offspring through germ cells. Jonathan Weissman and his lab have created a “map” which researchers can use to find the function of a given gene, allowing them access to an organism’s most fundamental plans. Read on to learn about these scientists’ work, and more.
Laying out the plan
All multicellular organisms begin with a single cell, the fertilized egg. This cell has an essential role in setting out the body plan for the rest of an organism. It all starts with establishing polarity — in other words, figuring out which side of the cell is the top, and which is the bottom. This polarity establishes an axis of symmetry for the growing organism, and sets the stage for other developmental processes to come.
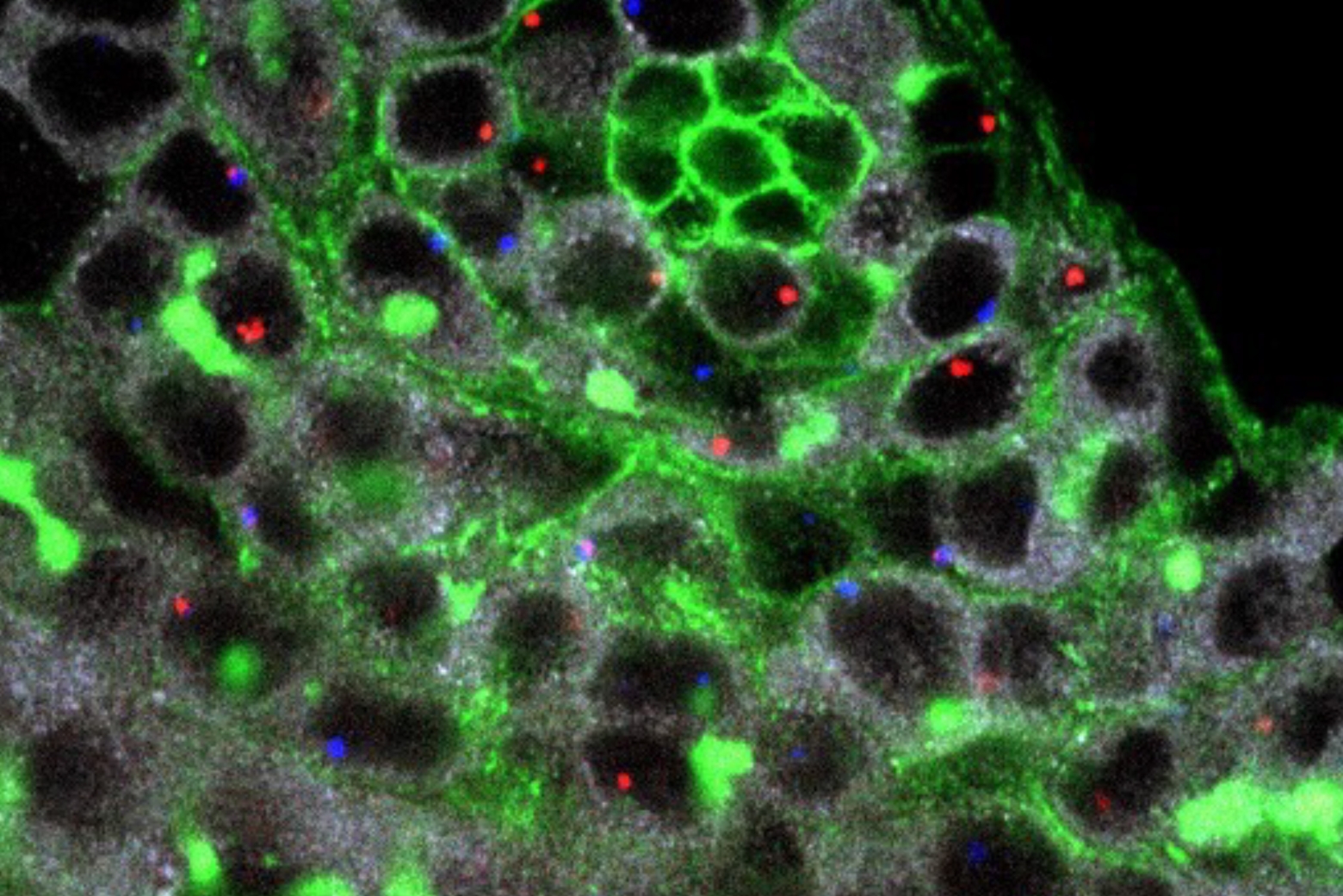
In a 2021 study, Cheeseman and postdoctoral researcher Zak Swartz investigated how one protein in specific, called Disheveled, localizes in a cell to help create this polarity in sea star embryos. Swartz found that Disheveled started out uniformly distributed in small aggregations throughout the egg cell, or oocyte. As the cell prepared to divide, Disheveled aggregations dissolved and then reformed at what would become the “bottom” of the oocyte.
Once the initial polarity is established, the oocyte can divide, creating a bilaterally symmetric sea star larvae. The burgeoning cluster of cells must then undergo other processes to define the several axes of symmetry that adult sea stars are known for.
Talking through it
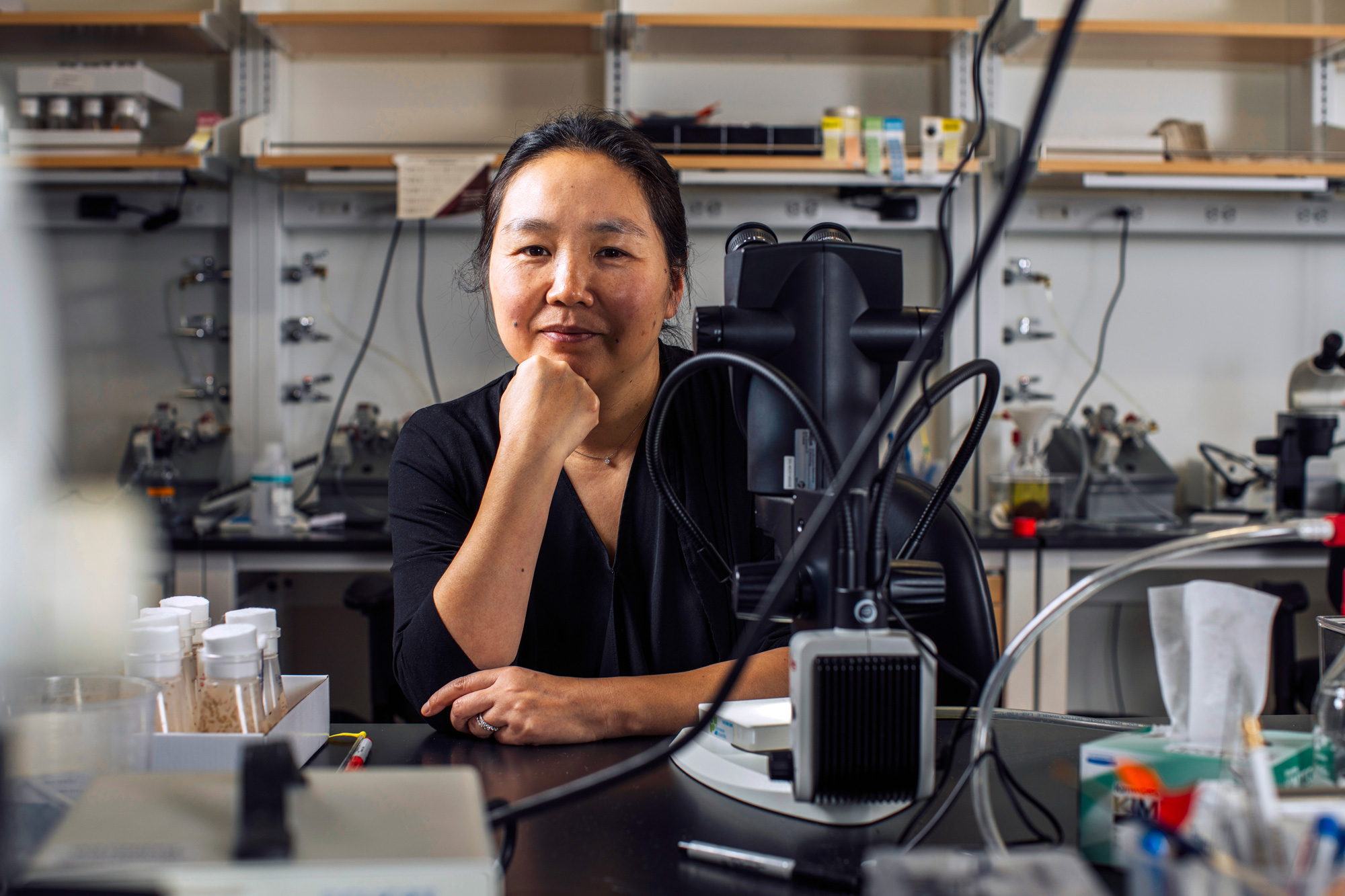
If an organism’s developmental blueprints are to be followed as development progresses, cells must be able to effectively communicate with each other. That cell to cell communication is the area of expertise of Whitehead Institute Member Pulin Li.
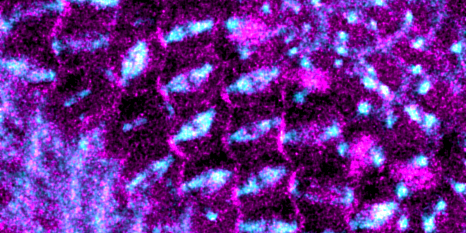
During her postdoctoral fellowship at the California Institute of Technology, Li studied tissue patterning — the mechanisms by which an organism’s newly forming tissues are laid out. Specifically, she investigated a developmental mechanism called morphogen gradient formation.
These gradients, composed of chemicals present in developing embryos, function as spatial coordinate systems and help determine how various cell types will be arranged in the organism — for example which groups of cells will form the liver, or the bones, or the brain, and where they will be within the body.
Li was able to recreate these gradients in the lab, in a Petri dish, and then interpret their signals using time lapse imaging and mathematical modeling. Here at Whitehead Institute, she follows a “bottom-up” approach to studying these complex systems. The best way to understand how something works, she says, is to build it yourself.