Graduate student Lauren Cote identifies genes directing regeneration
Justin Chen
November 16, 2017
An eye for a mouth: How regenerating flatworms keep track of body parts
Graduate student Lauren Cote identifies genes directing regeneration
Justin Chen
Peering down through a microscope at a petri dish, Lauren Cote, a sixth-year graduate student, watches the tip of a worm’s tail. Alone in the petri dish, the brown globule of tissue is regenerating an entirely new digestive system, a brain, and a pair of eye spots. After just a few weeks, the animal — a quarter-inch-long ribbon of flesh capped by a triangular head — is complete again. Swimming through the dish, the worm’s grainy, mahogany body fades to a translucent gray-blue along the edges, stretching and contracting as if hinting at its malleability.
Many animals regenerate. Salamanders replace their tails while zebrafish regrow damaged heart muscle. Even humans can renew large parts of their livers. However, few creatures can regenerate like planarians, a class of flatworms found in fresh and salt water habitats around the world — and in the Reddien lab at the Whitehead Institute.
Because planarians are masters of regeneration, able to replace any body part and even create a new animal from small chunks of tissue, they have become a focus of intense study. By examining the flatworm species Schmidtea mediterranea, Cote and other members of the Reddien lab have uncovered the ways cells communicate after injury to coordinate regeneration. Their work provides insight into how the ability to regenerate evolved, and how the healing process works in a variety of animals, including humans.
– –
Although regeneration seems mysterious, researchers have simplified the feat into two steps. First, planarians create the raw material to make new body parts by stimulating a group of rapidly dividing cells, called stem cells, that are the source of all new tissue in the worm. Second, these new cells need instructions to know what kind of tissue to become. Cote’s goal is to demystify this second step by locating a grid of information, like latitude and longitude lines on a map, that helps planarians keep track of their body parts and sense what is missing.
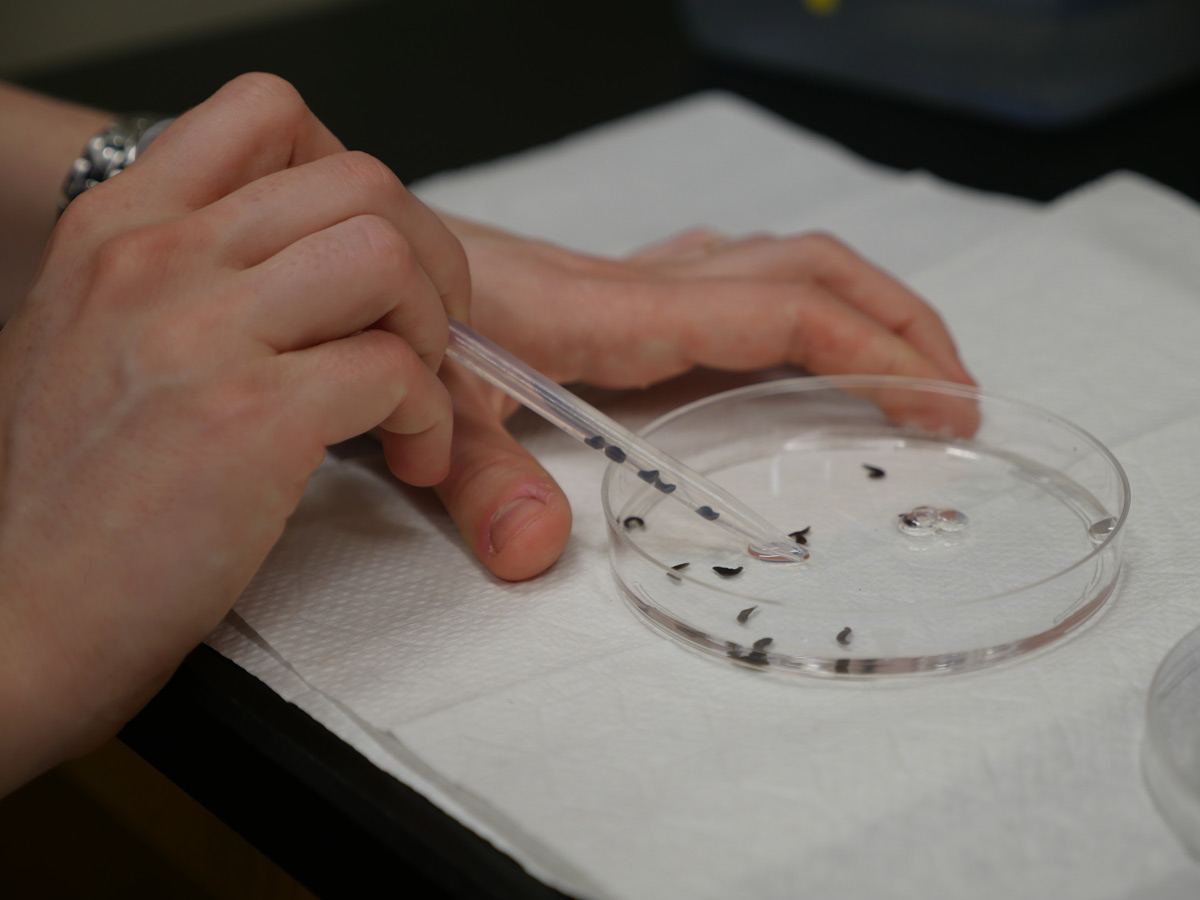

“The animal could have lost just the tip of its head or entire left side of its body,” Cote says, “and somehow it regrows the precise anatomy needed to make a complete worm.”
Over the past few years, research in the Reddien lab has demonstrated that a network of muscle cells spread throughout the worm’s body guides regeneration. To accomplish this task, muscle cells rely on a group of genes called position control genes (PCGs) which, based on Cote’s model, are predicted to encode proteins involved in cell communication. Depending on what PCGs are activated or expressed, muscle cells would send out a unique combination of signaling molecules that determine which body parts, such as eyes, stomach, or tail, would form.
“We like to imagine that muscle cells function like satellites and beam down information,” Cote says. “This allows stem cells to know where they are and what new body part to become.”
– –
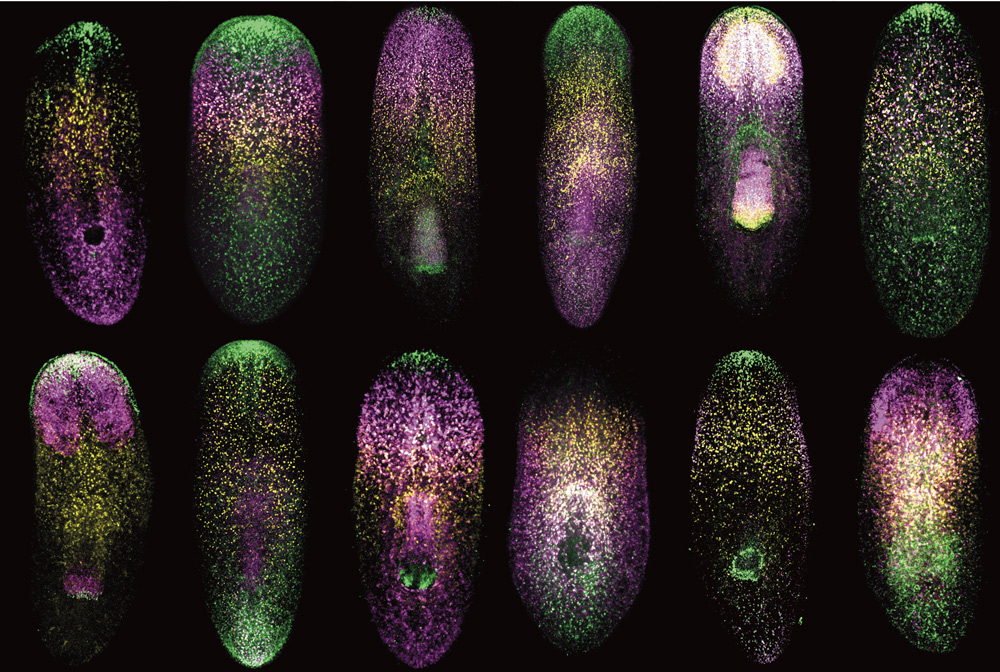
To systematically identify PCGs from the roughly 20,000 genes expressed in Schmidtea mediterranea, Cote worked in tandem with postdoctoral researcher Lucila Scimone in the Reddien lab to perform a two-part study. First they created maps of gene expression by examining individual muscle cells. After inventorying the genes each individual muscle cell expressed, they aggregated the data into a whole body map, showing gene activity across the entire worm. Some genes were expressed in all muscle cells, implying a general function such as controlling contraction and relaxation. In contrast, other genes were expressed in precise regions of the worm, like the head or midsection, suggesting that they could act as PCGs by defining the identity of each area.
In the second half of the study, Cote and Scimone used molecular techniques to disrupt the activity of potential PCGs. “We hypothesized that if a gene were needed to direct regeneration, the worm would still be able to renew itself without that gene’s activity,” Cote says, “but the animal would end up with an abnormal body.”
Indeed, Cote found that disrupting four genes in particular, encoding signaling molecules and receptor-like proteins, led to defective regeneration; worms either grew extra eyes on their head or grew extra feeding tubes sprouting out of their midsection like elongated suction cups. Together these four genes, along with a few previously identified genes controlling head and tail regeneration, comprise a short but expanding list of PCGs controlling the location and identity of new tissues. As scientists begin to understand the molecular details of planarian regeneration, they will test whether similar genes are used by other animals and humans.
– –
Although a biologist now, Cote began her academic life focusing on mathematics. As an undergraduate math major at the University of Chicago, she studied branches of mathematics such as analysis, algebra, and algebraic topology, a discipline that describes the properties of multidimensional shapes. After a summer project, Cote realized that — while she enjoyed learning mathematics — she found the research far too abstract.
“I was having a mid-college crisis,” she recalls. “I wanted to study something more visual where you could actually see what is going on.” Following this urge, Cote began to work in a lab examining fly development during her junior year. “I remember watching sheets of cells on the outside of a fly embryo folding in on themselves and sliding under the surface away from view. It made me wonder how cells make decisions and choreograph their movements to build a body. That’s how I got interested in developmental biology.”
After graduating from the University of Chicago, Cote worked as a lab technician for two years. During this time, she realized that her background in math and ability to think logically was an asset. “Putting together a mathematical proof is similar to publishing a research paper,” she says. “In both cases you are piecing together smaller bits of evidence into a cohesive argument.”
Encouraged by her successful venture into biological research, Cote decided to pursue a PhD in biology. She learned about the Reddien lab while taking a genetics course during her first year at MIT. Like Cote, many members of this group have backgrounds in other areas of science — including computational biology, development, evolution, biochemistry, and immunology — which helps them examine planarian regeneration from many perspectives.
“They were beginning to put together a story linking muscle cells to regeneration that was really intriguing,” Cote says. “I also liked the challenge of working with planarians because they are a fairly new lab animal. We’re still developing a lot of research tools so there is room to be creative and ask fundamental questions.”
By following an initial strand of curiosity as an undergraduate and identifying PCGs as a graduate student, Cote has begun to decipher the molecular language of regeneration. As scientists learn more about how planarians replace missing body parts, new areas of exploration open. One pressing question is how planarian regeneration compares to that of other animals. To pursue that mystery, Cote plans on studying another animal as a postdoctoral researcher and eventually starting her own laboratory.
“I still haven’t made up my mind, “she says, “but I’m considering a lot of possibilities such as crustaceans, sea squirts, zebrafish, and axolotls.” Regardless of her final choice, Cote will be investigating how cells — essentially fatty membranes encasing a slurry of water and proteins — manage to form complex and intricate structures. She will be pursuing the same questions that first captivated her as an undergraduate in Chicago. “How do cells make decisions? How do they know to become an eye or a stomach or a brain?” she asks. “There is a lot more that I want to understand.”