Greta Friar | Whitehead Institute
September 25, 2019
Cambridge, MA — During embryonic development, stem cells begin to take on specific identities, becoming distinct cell types with specialized characteristics and functions, in order to form the diverse organs and systems in our bodies. Cells rely on two main classes of regulators to define and maintain their identities; the first of these are master transcription factors, keystone proteins in each cell’s regulatory network, which keep the DNA sequences associated with crucial cell identity genes accessible for transcription — the process by which DNA is “read” into RNA. The other main regulators are signaling factors, which transmit information from the environment to the nucleus through a chain of proteins like a game of cellular telephone. Signaling factors can prompt changes in gene transcription as the cells react to that information.
One long-standing conundrum of how cell identity is determined is that many species, including humans, use the same core signaling pathways, with the same signaling factors, in all of their cells, yet this uniform machinery can cue a diverse array of cell-type specific gene activity, like an identical line of code being entered in many computers and causing each to start running a completely different program. New research from Whitehead Institute Member Richard Young, who is also a professor of biology at the Massachusetts Institute of Technology, published online in Molecular Cell on September 25, sheds light on how the same signaling factor can lead to so many distinct responses — with the help of a mechanism called phase separation.
Co-senior author on the paper Jurian Schuijers, previously a postdoctoral researcher in Young’s lab and now a professor at the Center for Molecular Medicine at the University Medical Center Utrecht, was drawn to this puzzle after previously working in a signaling lab: “Two cells of different types that are right next to each other in the body can receive the exact same signal and have different reactions, and there was not a satisfying explanation for how that happens,” Schuijers says.
Young’s lab had previously found that signaling factors in pathways important for development tend to concentrate at super enhancers, clusters of DNA sequences that increase transcription of crucial cell identity genes. Because super enhancers are established at the genes important to identity in each cell, their activity is cell-type specific, so this co-localization provided a partial explanation of the puzzle, but it raised the question of how signaling factors are recruited to super enhancers. Young found the vague explanations that had been put forward, such as super enhancers being the most accessible DNA to signaling factors and their co-factors, unconvincing.
Young and his team suspected that an explanation for signaling factor recruitment might lie in their research on transcriptional condensates — droplets that form at super enhancers and concentrate transcriptional machinery there using phase separation, meaning the molecules separate out of their surroundings to form a distinct liquid compartment, like a drop of vinegar in a pool of oil. The proteins in condensates can do this because they contain intrinsically disordered regions (IDRs), stretches of amino acids that remain flexible, like wet spaghetti, and do not become fixed into a single shape the way most protein structures do. This property allows them to mesh together to form a condensate. The researchers reasoned that if signaling factors were joining transcriptional condensates, that could explain their concentration at super enhancers.
Young’s team confirmed that signaling factors in several of the most important pathways for embryonic development in mammals — WNT, TGF-β and JAK/STAT — contained IDRs. They further found that these factors were able to use their IDRs to form and join condensates, and that, in mouse cells, they appeared to join the condensates at super enhancers upon activation of their respective pathways.
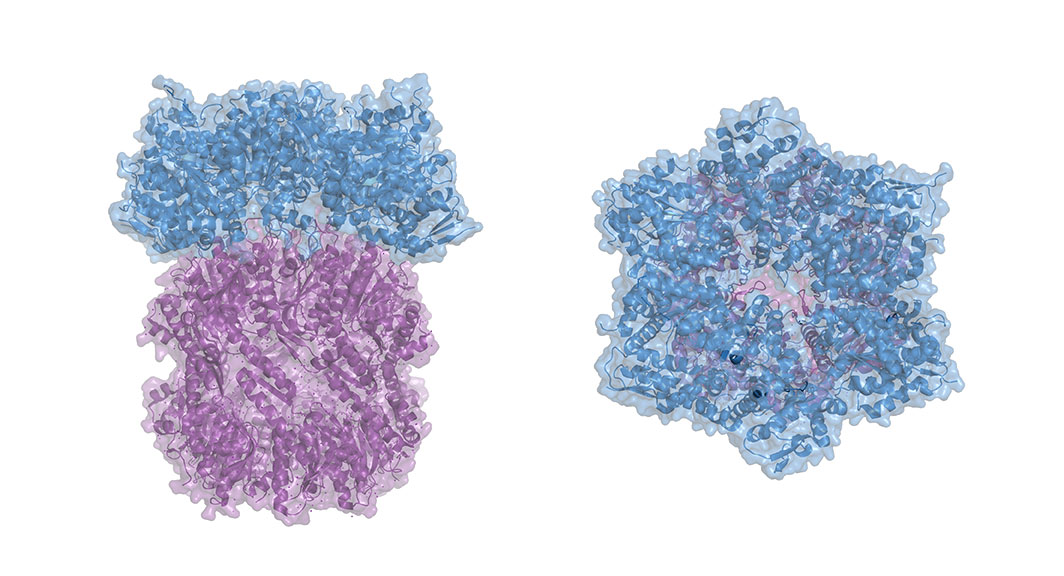
The researchers then decided to focus on beta catenin, the signaling factor at the end of the Wnt signaling pathway, a pathway essential for development; it helps to coordinate things like body axis patterning and cell fate specification, proliferation and migration. When Wnt signaling goes awry in embryos, they fail to develop, and when it goes awry in adults it is implicated in diseases including cancer. The beta catenin protein has IDRs on both of its ends and a structured middle section, called the Armadillo repeat domain, where it binds to other transcription factors. Typically, beta catenin binds to transcription factors in the TCF/LEF family, which in turn bind to DNA—beta catenin cannot bind to DNA on its own — anchoring the signaling factor at the right site and prompting gene transcription. However, the researchers found that beta catenin could concentrate at super enhancers even when it could not bind to its usual partners, suggesting that transcriptional condensates were a sufficient recruitment mechanism. The researchers then created two abridged beta catenin molecules: one version that only contained the IDRs and one that only contained the Armadillo repeat domain. Both partial factors were able to concentrate at super enhancers, but neither was as effective as the combined whole.
“If you ask most people how these factors find their target locations in the genome, they would say it’s through their DNA binding domains,” says first author Alicia Zamudio, a graduate student in Young’s lab. “This research suggests that factors use both their structured DNA binding domains and their unstructured domains to find the right locations to bind in the genome and to activate target genes.”
One advantage for cells of using IDRs, versus DNA binding alone, might be reducing the time it takes for signaling factors to concentrate near the right genes, the researchers say. Speed is of the essence for some signaling pathways in order for cells to be able to respond quickly to environmental stimuli. Transcriptional condensates are larger in size and much fewer in number than DNA binding sites or DNA-binding co-factors, and so they shrink the space that a signaling factor entering the nucleus must search.
This research could provide new opportunities for drug discovery. Signaling pathways and super enhancers are both co-opted by oncogenes to drive the spread of cancer, so transcriptional condensates could be a promising target to disrupt both oncogenic signaling and oncogene transcription. Young also hopes that this research, which adds to his lab’s growing body of work on transcriptional condensates, will lead to a new appreciation of the disordered regions of proteins.
“For a long time, researchers have mostly ignored the intrinsically disordered regions of proteins — we literally cut them off when identifying the crystal structures — much in the same way that researchers used to study genes and ignore ‘junk DNA,’” Young says. “But, just as with junk DNA, we are discovering that the overlooked, less obviously functional regions of these molecules are very important after all.”
This work is supported by NIH grant GM123511 and NSF grant PHY1743900 (R.A.Y.), NIH grant GM117370 (D.J.T.), NSF Graduate Research Fellowship (A.V.Z.), NIH grant T32CA009172 (I.A.K.), and DFG Research Fellowship DE 3069/1-1 (T.M.D.).
Written by Greta Friar
***
Richard Young’s primary affiliation is with Whitehead Institute for Biomedical Research, where his laboratory is located and all his research is conducted. He is also a professor of biology at the Massachusetts Institute of Technology.
***
Full citation:
“Mediator condensates localize signaling factors to key cell identity genes”
Molecular Cell, published online September 25, 2019. DOI: 10.1016/j.molcel.2019.08.016
Alicia V. Zamudio (1, 2), Alessandra Dall’Agnese (1), Jonathan E. Henninger (1), John C. Manteiga(1, 2), Lena K. Afeyan (1, 2), Nancy M. Hannett (1), Eliot L. Coffey (1, 2), Charles H. Li (1, 2), Ozgur Oksuz (1), Benjamin R. Sabari (1), Ann Boija (1), Isaac A. Klein (1,3), Susana W. Hawken (4), Jan-Hendrik Spille (5), Tim-Michael Decker (6), Ibrahim I. Cisse (5), Brian J. Abraham (1,7), Tong I. Lee (1), Dylan J. Taatjes (6), Jurian Schuijers (1,8,9), and Richard A. Young (1, 2, 9).
1. Whitehead Institute for Biomedical Research, Cambridge, MA 02142, USA
2. Department of Biology, Massachusetts Institute of Technology, Cambridge, MA, 02139, USA 3. Department of Medical Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA, 02215, USA
4. Program in Computational and Systems Biology, Massachusetts Institute of Technology, Cambridge, MA, 02139 USA
5. Department of Physics, Massachusetts Institute of Technology, Cambridge, MA, 02139, USA 6. Department of Biochemistry, University of Colorado, Boulder, CO 80303, USA
7. St. Jude Children’s Research Hospital, Memphis, TN, 038105, USA
8. Present address: Center for Molecular Medicine, University Medical Center Utrecht, Utrecht, 3584 CX, The Netherlands.
9. Equal contribution