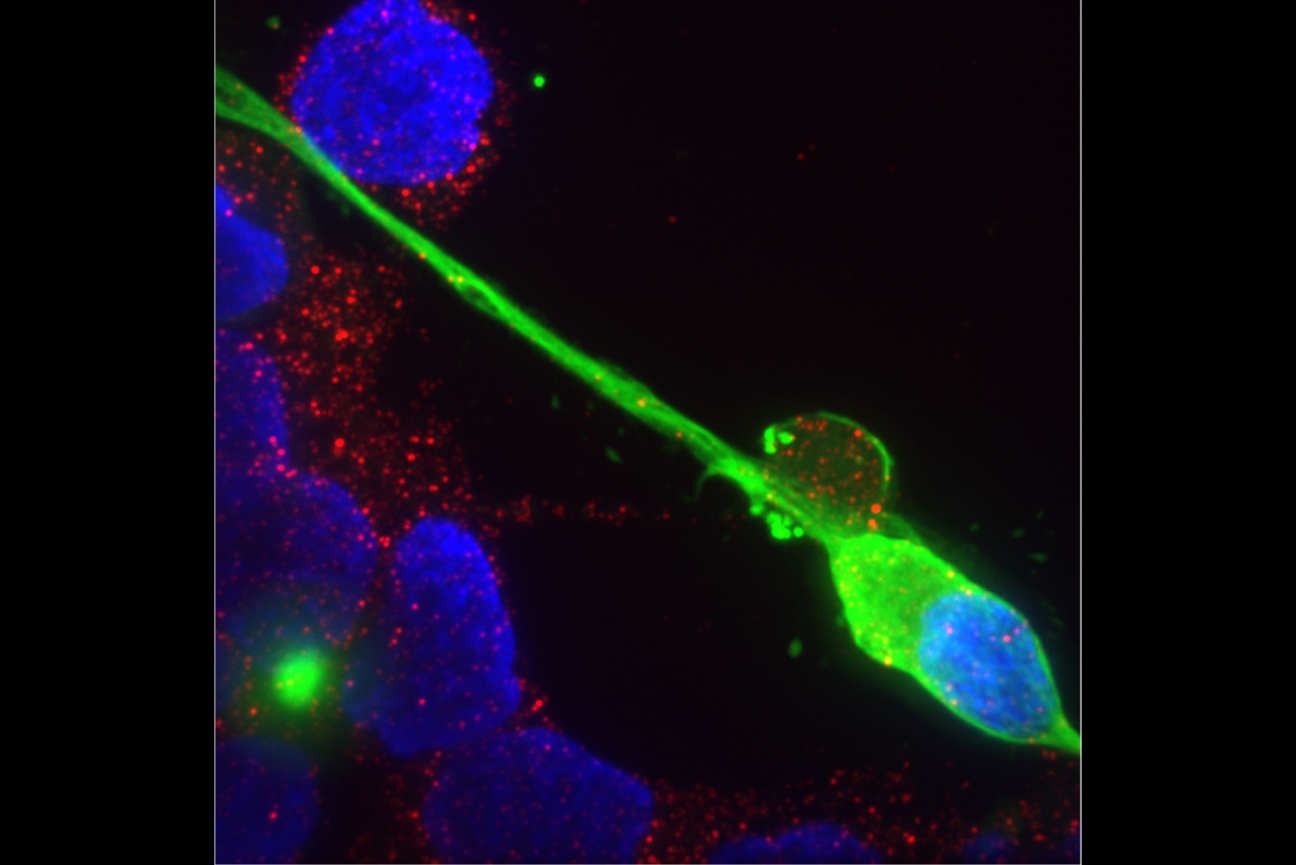



Sipping a beer on a warm summer evening, one might not consider that humans and yeast have been inextricably linked for thousands of years; winemaking, baking, and brewing all depend on budding yeast. Outside of baking and fermentation, researchers also use Saccharomyces cerevisiae, classified as a fungus, to study fundamental questions of cell biology.
Budding yeast gets its name from the way it multiplies. A daughter cell forms first as a swelling, protruding growth on the mother cell. The daughter cell projects further and further from the mother cell until it detaches as an independent yeast cell.
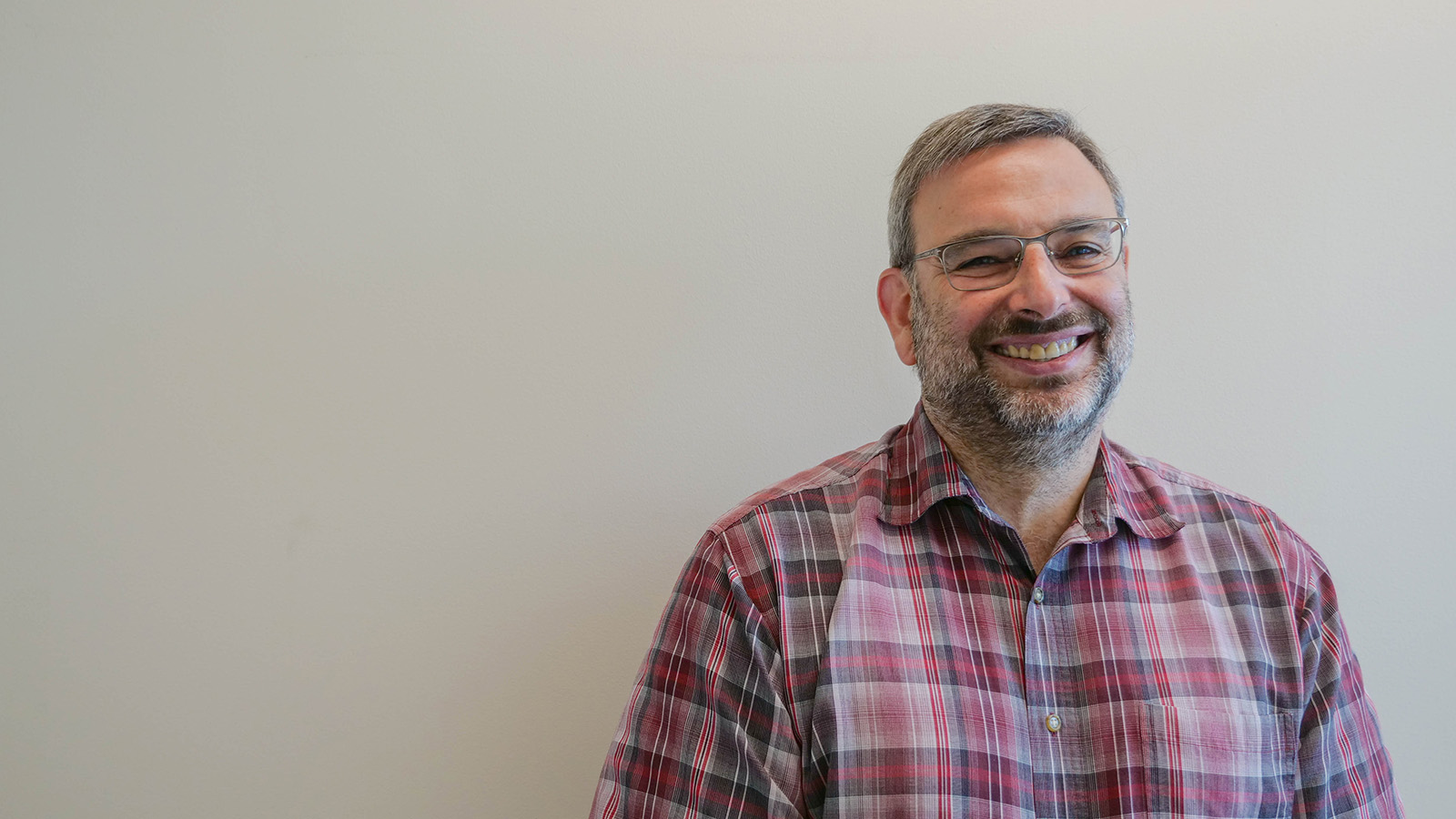
How do cells decide on a front and back? How do cells decode concentration gradients of chemical signals to orient in useful directions, or sense and navigate around physical obstacles? New Department of Biology faculty member Daniel “Danny” Lew uses the model yeast S. cerevisiae, and a non-model yeast with an unusual pattern of cell division, to explore these questions.
Q: Why is it useful to study yeast, and how do you approach the questions you hope to answer?
A: Humans and yeast are descended from a common ancestor, and some molecular mechanisms developed by that ancestor have been around for so long that yeast and mammals often use the same mechanisms. Many cells develop a front and migrate or grow in a particular direction, like the axons in our nervous system, using similar molecular mechanisms to those of yeast cells orienting growth towards the bud.
When I started my lab, I was working on cell cycle control, but I’ve always been interested in morphogenesis and the cell biology of how cells change shape and decide to do different things with different parts of themselves. Those mechanisms turn out to be conserved between yeast and humans.
But some things are very different about fungal and animal cells. One of the differences is the cell wall and what fungal cells have to do to deal with the fact that they have a cell wall.
Fungi are inflated by turgor pressure, which pushes their membranes against the rigid cell wall. This means they’ll die if there is any hole in the cell wall, which would be expected to happen often as cells remodel the wall in order to grow. We’re interested in understanding how fungi sense when any weak spots appear in the wall and repair them before those weak spots become dangerous.
Yeast cells, like most fungi, also mate by fusing with a partner. To succeed, they must do the most dangerous thing in the fungal lifecycle: get rid of the cell wall at the point of contact to allow fusion. That means they must be precise about where and when they remove the wall. We’re fascinated to understand how they know it is safe to remove the wall there, and nowhere else.
We take an interdisciplinary approach. We’ve used genetics, biochemistry, cell biology, and computational biology to try and solve problems in the past. There’s a natural progression: observation and genetic approaches tend to be the first line of attack when you know nothing about how something works. As you learn more, you need biochemical approaches and, eventually, computational approaches to understand exactly what mechanism you’re looking at.
I’m also passionate about mentoring, and I love working with trainees and getting them fascinated by the same problems that fascinate me. I’m looking to work with curious trainees who love addressing fundamental problems.
Q: How does yeast decide to orient a certain way—towards a mating partner, for example?
A: We are still working on questions of how cells analyze the surrounding environment to pick a direction. Yeast cells have receptors that sense pheromones that a mating partner releases. What is amazing about that is that these cells are incredibly small, and pheromones are released by several potential partners in the neighborhood. That means yeast cells must interpret a very confusing landscape of pheromone concentrations. It’s not apparent how they manage to orient accurately toward a single partner.
That got me interested in related questions. Suppose the cell is oriented toward something that isn’t a mating partner. The cell seems to recognize that there’s an obstacle in the way, and it can change direction to go around that obstacle. This is how fungi get so good at growing into things that look very solid, like wood, and some fungi can even penetrate Kevlar vests.
If they recognize an obstacle, they have to change directions and go around it. If they recognize a mating partner, they have to stick with that direction and allow the cell wall to get degraded. How do they know they’ve hit an obstacle? How do they know a mating partner is different from an obstacle? These are the questions we’d like to understand.
Q: For the last couple of years, you’ve also been studying a budding yeast that forms multiple buds when it reproduces instead of just one. How did you come across it, and what questions are you hoping to explore?
A: I spent several years trying to figure out why most yeasts make one bud and only one bud, which I think is related to the question of why migrating cells make one and only one front. We had what we thought was a persuasive answer to that, so seeing a yeast completely disobey that and make as many buds as it felt like was a shock, which got me intrigued.
We started working on it because my colleague,Amy Gladfelter, had sampled the waters around Woods Hole, Massachusetts. When she saw this specimen under a microscope, she immediately called me and said, “You have to look at this.”
A question we’re very intrigued by is if the cell makes five, seven, or 12 buds simultaneously, how do they divide the mother cell’s material and growth capacity five, seven, or 12 ways? It looks like all of the buds grow at the same rate and reach about the same size. One of our short-term goals is to check whether all the buds really get to exactly the same size or whether they are born unequal.
And we’re interested in more than just growth rate. What about organelles? Do you give each bud the same number of mitochondria, nuclei, peroxisomes, and vacuoles? That question will inevitably lead to follow-up questions. If each bud has the same number of mitochondria, how does the cell measure mitochondrial inheritance to do that? If they don’t have the same amount, then buds are each born with a different complement and ratio of organelles. What happens to buds if they have very different numbers of organelles?
As far as we can tell, every bud gets at least one nucleus. How the cell ensures that each bud gets a nucleus is a question we’d also very much like to understand.
We have molecular candidates because we know a lot about how model yeasts deliver nuclei, organelles, and growth materials from the mother to the single bud. We can mutate candidate genes and see if similar molecular pathways are involved in the multi-budding yeast and, if so, how they are working.
It turns out that this unconventional yeast has yet to be studied from the point of view of basic cell biology. The other thing that intrigues me is that it’s a poly-extremophile. This yeast can survive under many rather harsh conditions: it’s been isolated in Antarctica, from jet engines, from all kinds of plants, and of course from the ocean as well. An advantage of working with something so ubiquitous is we already know it’s not toxic to us under almost any circumstances. We come into contact with it all the time. If we learn enough about its cell biology to begin to manipulate it, then there are many potential applications, from human health to agriculture.
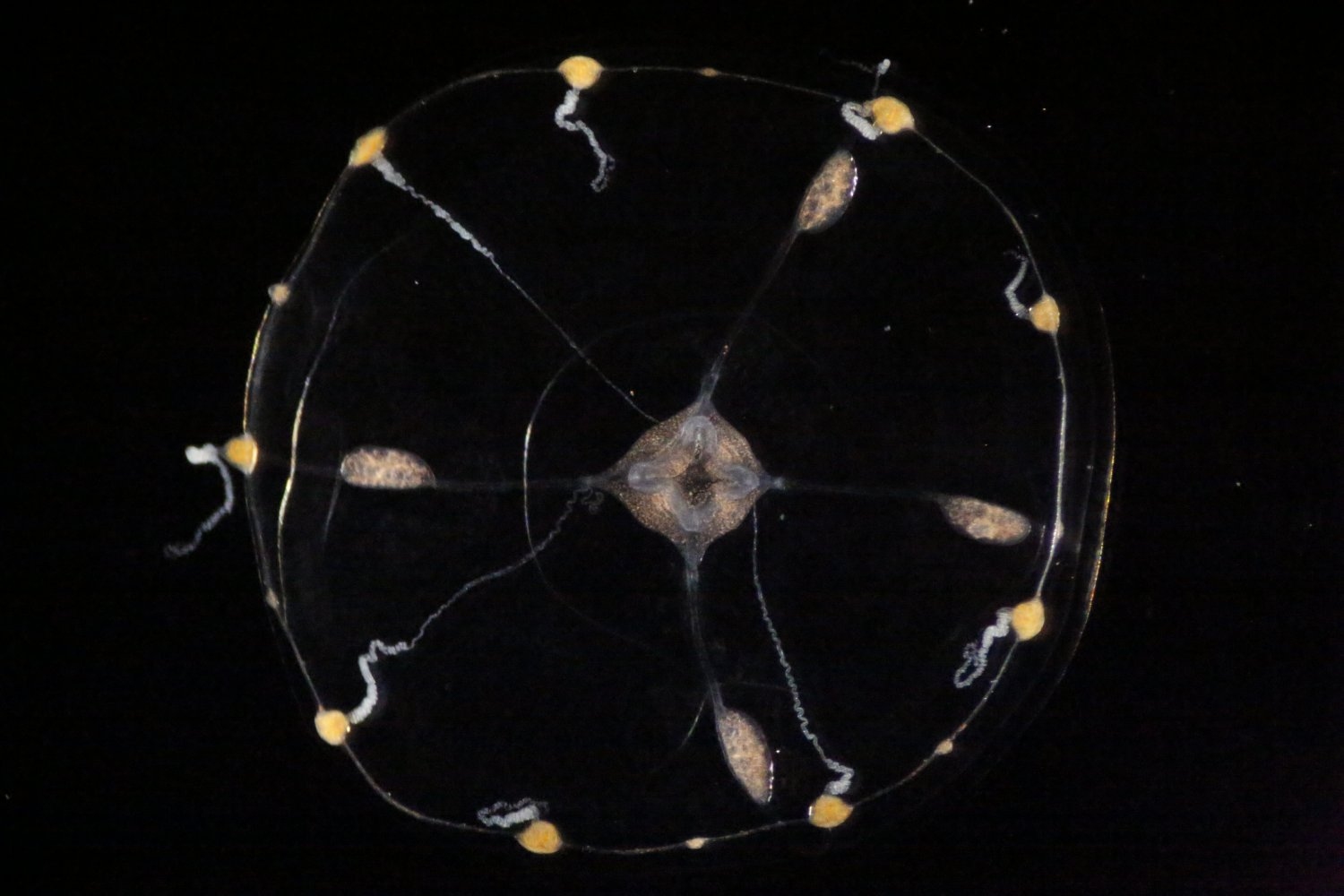
The Clytia hemisphaerica jellyfish is not only a hypnotically graceful swimmer, but also an amazing neuron-manufacturing machine with a remarkable ability to expand and regenerate its nervous system.
Now, thanks to a prestigious Klingenstein-Simons Fellowship Award in Neuroscience, MIT Assistant Professor Brady Weissbourd will study how the tiny, transparent animals use this ability to build, organize, and rebuild a stable, functional, and robust nervous system throughout their lives.
“As we look more broadly across the animal kingdom it is amazing to see how similar the basic biology is of animals that look completely different — even jellyfish have neurons similar to our own that generate their behavior,” says Weissbourd, a faculty member in MIT’s Department of Biology and The Picower Institute for Learning and Memory, whose work to engineer genetic access to C. hemisphaerica in 2021 established it as a new neuroscience model organism. “At the same time, it could be just as important to examine what is different across species, particularly when it comes to some of the incredible capabilities that have evolved.”
Weissbourd is just one of 13 researchers nationally to be recognized with this fellowship, which provides $300,000 over three years. It will enable Weissbourd’s lab to tackle several questions raised by the jellyfish’s prodigious production of neurons. Where does the constant stream of newborn neurons come from, and what guides them to their eventual places in the jellyfish’s mesh-like neural network? How does the jellyfish organize these ever-changing neural populations — for instance, into functional circuits — to enable its various behaviors?
Another question hails from the surprising results of an experiment in which Weissbourd ablated the entire class of the neurons that the jellyfish uses to fold up its umbrella-shaped body — about 10 percent of the 10,000 or so neurons that it has. He found that within a week enough new neurons had taken their place that the folding behavior was restored. Weissbourd’s studies will also seek to determine how the animal can so readily bounce back from the destruction of a whole major neural network and the behavior it produces.
“We were studying the neural control of a particular behavior and stumbled across this shocking observation that the subnetwork that controls this behavior is constantly changing size and can completely regenerate,” Weissbourd says. “We want to understand the mechanisms that allow this network to be so robust, including the ability to rebuild itself from scratch. I’m very grateful to the Klingenstein Fund and the Simons Foundation for supporting our work.”
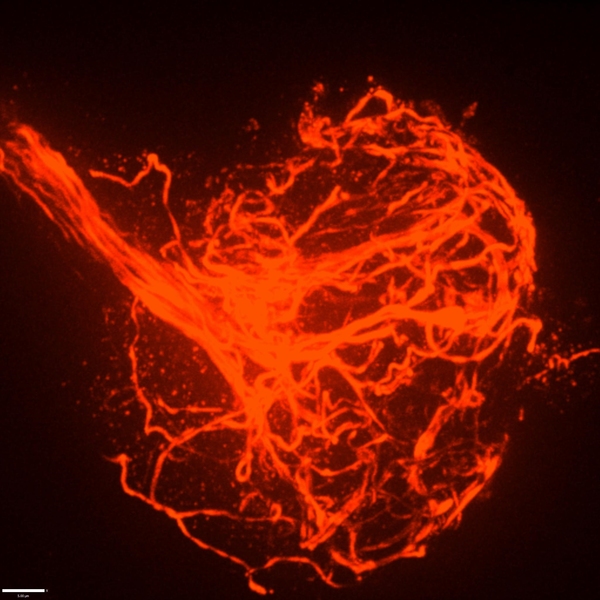
Perhaps the most obvious feature of a neuron is the long branch called an axon that ventures far from the cell body to connect with other neurons or muscles. If that long, thin projection ever seems like it could be vulnerable, a new MIT study shows that its structural integrity may indeed require the support of a surrounding protein called perlecan. Without that protein in Drosophila fruit flies, researchers at The Picower Institute for Learning and Memory found axonal segments can break apart during development and the connections, or synapses, that they form end up dying away.
Perlecan helps make the extracellular matrix, the proteins and other molecules that surround cells, stable and flexible so that cells can develop and function in an environment that is supportive without being rigid.
“What we found was that the extracellular matrix around nerves was being altered and essentially causing the nerves to break completely. Broken nerves eventually led to the synapses retracting,” says study senior author Troy Littleton, the Menicon Professor in MIT’s departments of Biology and Brain and Cognitive Sciences.
Humans need at least some perlecan to survive after birth. Mutations that reduce, but don’t eliminate, perlecan can cause Schwartz-Jampel syndrome, in which patients experience neuromuscular problems and skeletal abnormalities. The new study may help explain how neurons are affected in the condition, Littleton says, and also deepen scientists’ understanding of how the extracellular matrix supports axon and neural circuit development.
Ellen Guss PhD ’23, who recently defended her doctoral thesis on the work, led the research published June 8 in eLife.
At first she and Littleton didn’t expect the study to yield a new discovery about the durability of developing axons. Instead, they were investigating a hypothesis that perlecan might help organize some of the protein components in synapses that fly nerves develop to connect with muscles. But when they knocked out the gene called “trol” that encodes perlecan in flies, they saw that the neurons appeared to “retract” many synapses at a late stage of larval development. Proteins on the muscle side of the synaptic connection remained, but the neuron side of the connection withered away. That suggested that perlecan had a bigger role than they first thought.
Indeed, the authors found that the perlecan wasn’t particularly enriched around synapses. Where it was pronounced was in a structure called the neural lamella, which surrounds axon bundles and acts a bit like the rubbery cladding around a TV cable to keep the structure intact. That suggested that a lack of perlecan might not be a problem at the synapse, but instead causes trouble along axons due to its absence in the extracellular matrix surrounding nerve bundles.
Littleton’s lab had developed a technique for daily imaging of fly neural development called serial intravital imaging. They applied it to watch what happened to the fly axons and synapses over a four-day span. They observed that while fly axons and synapses developed normally at first, not only synapses but also whole segments of axons faded away.
They also saw that the farther an axon segment was from the fly’s brain, the more likely it was to break apart, suggesting that the axon segments became more vulnerable the further out they extended. Looking segment by segment, they found that where axons were breaking down, synapse loss would soon follow, suggesting that axon breakage was the cause of the synapse retraction.
“The breakages were happening in a segment-wide manner,” Littleton says. “In some segments the nerves would break and in some they wouldn’t. Whenever there was a breakage event, you would see all the neuromuscular junctions (synapses) across all the muscles in that segment retract.”
When they compared the structure of the lamella in mutant versus healthy flies, they found that the lamella was thinner and defective in the mutants. Moreover, where the lamella was weakened, axons were prone to break and the microtubule structures that run the length of the axon would become misdirected, protruding outward and becoming tangled up in dramatic bundles at sites of severed axons.
In one other key finding, the team showed that perlecan’s critical role depended on its secretion from many cells, not just neurons. Blocking the protein in just one cell type or another did not cause the problems that total knockdown did, and enhancing secretion from just neurons was not enough to overcome its deficiency from other sources.
Altogether, the evidence pointed to a scenario where lack of perlecan secretion caused the neural lamella to be thin and defective, with the extracellular matrix becoming too rigid. The further from the brain nerve bundles extended, the more likely movement stresses would cause the axons to break where the lamella had broken down. The microtubule structure within the axons then became disorganized. That ultimately led to synapses downstream of those breakages dying away because the disruption of the microtubules means the cells could no longer support the synapses.
“When you don’t have that flexibility, although the extracellular matrix is still there, it becomes very rigid and tight and that basically leads to this breakage as the animal moves and pulls on those nerves over time,” Littleton says. “It argues that the extracellular matrix is functional early on and can support development, but doesn’t have the right properties to sustain some key functions over time as the animal begins to move and navigate around. The loss of flexibility becomes really critical.”
In addition to Littleton and Guss, the paper’s other authors are Yulia Akbergenova and Karen Cunningham.
Support for the study came from the National Institutes of Health. The Littleton Lab is also supported by The Picower Institute for Learning and Memory and The JPB Foundation.

Humans split away from our closest animal relatives, chimpanzees, and formed our own branch on the evolutionary tree about seven million years ago. In the time since—brief, from an evolutionary perspective—our ancestors evolved the traits that make us human, including a much bigger brain than chimpanzees and bodies that are better suited to walking on two feet. These physical differences are underpinned by subtle changes at the level of our DNA. However, it can be hard to tell which of the many small genetic differences between us and chimps have been significant to our evolution.
New research from Whitehead Institute Member Jonathan Weissman; University of California, San Francisco Assistant Professor Alex Pollen; Weissman lab postdoc Richard She; Pollen lab graduate student Tyler Fair; and colleagues uses cutting edge tools developed in the Weissman lab to narrow in on the key differences in how humans and chimps rely on certain genes. Their findings, published in the journal Cell on June 20, may provide unique clues into how humans and chimps have evolved, including how humans became able to grow comparatively large brains.
Studying function rather than genetic code
Only a handful of genes are fundamentally different between humans and chimps; the rest of the two species’ genes are typically nearly identical. Differences between the species often come down to when and how cells use those nearly identical genes. However, only some of the many differences in gene use between the two species underlie big changes in physical traits. The researchers developed an approach to narrow in on these impactful differences.
Their approach, using stem cells derived from human and chimp skin samples, relies on a tool called CRISPR interference (CRISPRi) that Weissman’s lab developed. CRISPRi uses a modified version of the CRISPR/Cas9 gene editing system to effectively turn off individual genes. The researchers used CRISPRi to turn off each gene one at a time in a group of human stem cells and a group of chimp stem cells. Then they looked to see whether or not the cells multiplied at their normal rate. If the cells stopped multiplying as quickly or stopped altogether, then the gene that had been turned off was considered essential: a gene that the cells need to be active–producing a protein product–in order to thrive. The researchers looked for instances in which a gene was essential in one species but not the other as a way of exploring if and how there were fundamental differences in the basic ways that human and chimp cells function.
By looking for differences in how cells function with particular genes disabled, rather than looking at differences in the DNA sequence or expression of genes, the approach ignores differences that do not appear to impact cells. If a difference in gene use between species has a large, measurable effect at the level of the cell, this likely reflects a meaningful difference between the species at a larger physical scale, and so the genes identified in this way are likely to be relevant to the distinguishing features that have emerged over human and chimp evolution.
“The problem with looking at expression changes or changes in DNA sequences is that there are many of them and their functional importance is unclear,” says Weissman, who is also a professor of biology at the Massachusetts Institute of Technology and an Investigator with the Howard Hughes Medical Institute. “This approach looks at changes in how genes interact to perform key biological processes, and what we see by doing that is that, even on the short timescale of human evolution, there has been fundamental rewiring of cells.”
After the CRISPRi experiments were completed, She compiled a list of the genes that appeared to be essential in one species but not the other. Then he looked for patterns. Many of the 75 genes identified by the experiments clustered together in the same pathways, meaning the clusters were involved in the same biological processes. This is what the researchers hoped to see. Individual small changes in gene use may not have much of an effect, but when those changes accumulate in the same biological pathway or process, collectively they can cause a substantive change in the species. When the researchers’ approach identified genes that cluster in the same processes, this suggested to them that their approach had worked and that the genes were likely involved in human and chimp evolution.
“Isolating the genetic changes that made us human has been compared to searching for needles in a haystack because there are millions of genetic differences, and most are likely to have negligible effects on traits,” Pollen says. “However, we know that there are lots of small effect mutations that in aggregate may account for many species differences. This new approach allows us to study these aggregate effects, enabling us to weigh the impact of the haystack on cellular functions.”
Researchers think bigger brains may rely on genes regulating how quickly cells divide
One cluster on the list stood out to the researchers: a group of genes essential to chimps, but not to humans, that help to control the cell cycle, which regulates when and how cells decide to divide. Cell cycle regulation has long been hypothesized to play a role in the evolution of humans’ large brains. The hypothesis goes like this: Neural progenitors are the cells that will become neurons and other brain cells. Before becoming mature brain cells, neural progenitors divide multiple times to make more of themselves. The more divisions that the neural progenitors undergo, the more cells the brain will ultimately contain—and so, the bigger it will be. Researchers think that something changed during human evolution to allow neural progenitors to spend less time in a non-dividing phase of the cell cycle and transition more quickly towards division. This simple difference would lead to additional divisions, each of which could essentially double the final number of brain cells.
Consistent with the popular hypothesis that human neural progenitors may undergo more divisions, resulting in a larger brain, the researchers found that several genes that help cells to transition more quickly through the cell cycle are essential in chimp neural progenitor cells but not in human cells. When chimp neural progenitor cells lose these genes, they linger in a non-dividing phase, but when human cells lose them, they keep cycling and dividing. These findings suggest that human neural progenitors may be better able to withstand stresses—such as the loss of cell cycle genes—that would limit the number of divisions the cells undergo, enabling humans to produce enough cells to build a larger brain.
“This hypothesis has been around for a long time, and I think our study is among the first to show that there is in fact a species difference in how the cell cycle is regulated in neural progenitors,” She says. “We had no idea going in which genes our approach would highlight, and it was really exciting when we saw that one of our strongest findings matched and expanded on this existing hypothesis.”
More subjects lead to more robust results
Research comparing chimps to humans often uses samples from only one or two individuals from each species, but this study used samples from six humans and six chimps. By making sure that the patterns they observed were consistent across multiple individuals of each species, the researchers could avoid mistaking the naturally occurring genetic variation between individuals as representative of the whole species. This allowed them to be confident that the differences they identified were truly differences between species.
The researchers also compared their findings for chimps and humans to orangutans, which split from the other species earlier in our shared evolutionary history. This allowed them to figure out where on the evolutionary tree a change in gene use most likely occurred. If a gene is essential in both chimps and orangutans, then it was likely essential in the shared ancestor of all three species; it’s more likely for a particular difference to have evolved once, in a common ancestor, than to have evolved independently multiple times. If the same gene is no longer essential in humans, then its role most likely shifted after humans split from chimps. Using this system, the researchers showed that the changes in cell cycle regulation occurred during human evolution, consistent with the proposal that they contributed to the expansion of the brain in humans.
The researchers hope that their work not only improves our understanding of human and chimp evolution, but also demonstrates the strength of the CRISPRi approach for studying human evolution and other areas of human biology. Researchers in the Weissman and Pollen labs are now using the approach to better understand human diseases—looking for the subtle differences in gene use that may underlie important traits such as whether someone is at risk of developing a disease, or how they will respond to a medication. The researchers anticipate that their approach will enable them to sort through many small genetic differences between people to narrow in on impactful ones underlying traits in health and disease, just as the approach enabled them to narrow in on the evolutionary changes that helped make us human.
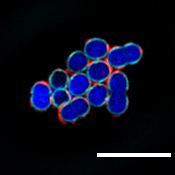
Your mouth is a crucial interface between the outside world and the inside of your body. Everything you breathe, chew or drink interacts with your oral cavity—the proteins and the microbes, including microbes that can harm us. When things go awry, the result can range from the mild, like bad breath, to the serious, like tooth and gum decay to more dire effects in the gut and other parts of the body.
Even though the oral microbiome plays a critical role as a front-line defense for human health and disease, we still know very little about the intricacies of host-microbe interactions in the complex physiological environment of the mouth; a better understanding of those interactions is key to developing treatments for human disease.
In a recent study published in PNAS, a collaborative effort revealed that one of the most abundant proteins found in our saliva binds to the surface of select microbes found in the mouth. The findings shed light on how salivary proteins and mucus play a role in maintaining the oral cavity microbiome.
The collaboration involved members of the Imperiali lab in the Department of Biology and the Kiessling lab in the Department of Chemistry at MIT, as well as the Ruhl group at the University at Buffalo School of Dental Medicine, and the Grimes group at the University of Delaware.
The paper is focused on an abundant oral cavity protein called zymogen granule protein 16 homolog B (ZG16B). Finding ZG16B’s interaction partners and gaining insight into its function were the overarching goals of the project. To accomplish this, Ghosh and colleagues engineered ZG16B to add reporter tags such as fluorophores. They called these modified proteins “microbial glycan analysis probes (mGAPs)” because they allowed them to identify ZG16B binding partners using complementary methods. They applied the probes to samples of healthy oral microbiomes to identify target microbes and binding partners.
The results excited them.
“ZG16B didn’t just bind to random bacteria. It was very focused on certain species including a commensal bacteria called Streptococcus vestibularis,” says first author Soumi Ghosh, a postdoctoral associate in the Imperiali lab.
Commensal bacteria are found in a normal healthy microbiome and do not cause disease.
Using the mGAPs, the team showed that ZG16B binds to cell wall polysaccharides of the bacteria, which indicates that ZG16B is a lectin, a carbohydrate-binding protein. In general, lectins are responsible for cell-cell interactions, signaling pathways, and some innate immune responses against pathogens. “This is the first time that it has been proven experimentally that ZG16B acts as a lectin because it binds to the carbohydrates on the cell surface or cell wall of the bacteria,” Ghosh highlights.
ZG16B was also shown to recruit Mucin 7 (MUC7), a salivary glycoprotein in the oral cavity, and, together the results suggest that ZG16B may help maintain a healthy balance in the oral microbiome by forming a complex with MUC7 and certain bacteria. The results indicate that ZG16B regulates the bacteria’s abundance by preventing overgrowth through agglutination when the bacteria exceed a certain level of growth.
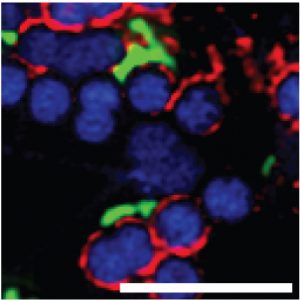
“ZG16B, therefore, seems to function as a missing link in the system; it binds to different types of glycans—the microbial glycans and the mucin glycans—and ultimately, maintains a healthy balance in our oral cavity,” Ghosh says.
Further work with this probe and samples of oral microbiome from healthy and diseased subjects could also reveal the lectin’s importance for oral health and disease.
Current attention is focused on developing and applying additional mGAPs based on other human lectins, such as those found in serum, liver, and intestine to reveal their binding specificities and their roles in host-microbe interactions.
“The research carried out in this collaboration exemplifies the kind of synergy that made me excited to move to MIT 5 years ago,” says senior co-author Laura Kiessling. “I’ve been able to work with outstanding scientists who share my interest in the chemistry and the biology of carbohydrates.”
The senior authors of the paper—Barbara Imperiali and Kiessling — came up with the term for the probes they’re creating: “mGAPS to fill in the gaps” in our understanding of the role of lectins in the human microbiome, according to Ghosh.
“If we want to develop therapeutics against bacterial infection, we need a better understanding of host-microbe interactions,” Ghosh says. “The significance of our study is to prove that we can make very good probes for microbial glycans, find out their importance in the frontline defense of the immune system, and, ultimately, come up with a therapeutic approach to disease.”
This research was supported by the National Institute of Health.

The National Academy of Sciences has elected 120 members and 23 international members, including five faculty members from MIT. Joshua Angrist, Gang Chen, Catherine Drennan, Dina Katabi, and Gregory Stephanopoulos were elected in recognition of their “distinguished and continuing achievements in original research.” Membership to the National Academy of Sciences is one of the highest honors a scientist can receive in their career.
Established in 1863 by a Congressional charter that was signed by Abraham Lincoln, the National Academy of Sciences is a private, nonprofit society of distinguished scholars. Each year, new members are elected by their peers in recognition of their outstanding contributions to their field of research. Together with the National Academy of Engineering and National Academy of Medicine, the National Academy of Sciences aims to “encourage education and research, recognize outstanding contributions to knowledge, and increase public understanding in matters of science, engineering, and medicine.”
As of this year, the National Academy of Sciences has 2,565 active members and 526 international members. Among the new members added this year are eight MIT alumni, including Thomas Banks PhD ’73; Joan W. Bresnan PhD ’72; Jennifer Elisseeff PhD ’99; current faculty member Dina Katabi SM ’99, PhD ’03; Maria C. Lemos SM ’90, PhD ’95; William B. McKinnon ’76; Emmanuel Saez PhD ’99; and Gunther Uhlmann PhD ’76.
Joshua Angrist
Joshua Angrist is the Ford Professor of Economics at MIT, a co-founder and director of MIT’s Blueprint Labs, and a research associate at the National Bureau of Economic Research. Angrist and his collaborators have pioneered the use of natural experiments to answer important economic questions and developed new econometric tools that help social scientists and policymakers discover the causal effects of individual choices and government policy changes. Angrist’s research explores the economics of education and school reform, the impact of social programs on the labor market, and the labor market effects of immigration, regulation, and economic institutions.
Angrist received his bachelor’s degree in economics from Oberlin College in 1982 and completed his PhD in economics at Princeton University in 1989. He taught at Harvard University and the Hebrew University of Jerusalem before coming to MIT in 1996.
Angrist received the Sveriges Riksbank Prize in Economic Sciences in Memory of Alfred Nobel in 2021 with co-laureates Guido Imbens of the Stanford Graduate School of Business and David Card of the University of California at Berkeley. Angrist is a fellow of the American Academy of Arts and Sciences and the Econometric Society, a Margaret MacVicar Faculty Fellow, and has served as co-editor of the Journal of Labor Economics.
Gang Chen
Gang Chen is the Carl Richard Soderberg Professor of Power Engineering in the Department of Mechanical Engineering. Chen is a pioneer in nanoscale heat transfer and energy conversion. He has significantly contributed to the understanding of heat transfer and energy conversion mechanisms; developed high-performance thermoelectric materials, superior semiconductors, highly heat-conductive polymers, and water desalination materials; and advanced solar-thermal and solar photovoltaic technologies. Physics World chose Chen’s work on cubic boron arsenide being a superior semiconductor as a Top 10 Breakthrough in 2022. Scientific American highlighted his directional solvent extraction and thermally charged batteries technologies as one of its annual top 10 World Changing Ideas in 2012 and 2014. His work on high-performance thermoelectric materials won an R&D 100 award.
Chen earned both his bachelor’s and master’s degrees from Huazhong University of Science and Technology in China and his PhD from UC Berkeley. He worked at Duke University and UCLA before joining the MIT faculty in 2001. Chen served as department head of MIT’s Department of Mechanical Engineering from 2013 to 2018 and director of the Solid-State Solar-Thermal Energy Conversion Center for the U.S. Department of Energy EFRC from 2009 to 2018.
Chen is a dedicated mentor and advocate for diversity and inclusion in STEM fields. He has supervised 86 master’s and PhD theses and 60 postdocs. Chen received an NSF Young Investigator Award, an ASME Heat Transfer Memorial Award, an ASME Frank Kreith Award in Energy, a Nukiyama Memorial Award from Japan Heat Transfer Society, a World Technology Network Award in Energy, an SES Eringen Medal, and the Capers and Marion McDonald Award for Excellence in Mentoring and Advising from MIT. He is an academician of Academy Sinica, a fellow of the American Academy of Arts and Sciences, and a National Academy of Engineering member.
Catherine Drennan
Catherine Drennan, professor of biology and chemistry, combines X-ray crystallography, cryo-electron microscopy and other biophysical methods, with the goal of “visualizing” molecular processes by obtaining snapshots of enzymes in action.
Drennan earned her bachelor’s degree from Vassar College, and her PhD from the University of Michigan. Following a postdoctoral fellowship at Caltech, she joined the MIT faculty in 1999, and was named a Howard Hughes Medical Institute Professor in recognition of her teaching in 2006 and a Howard Hughes Medical Institute Investigator in recognition of her research in 2008. Drennan has led by example, dedicating herself to both research and teaching. Her educational initiatives include creating free resources for educators that help students recognize the underlying chemical principles in biology and medicine, and training graduate student teaching assistants and mentors to be effective teacher–scholars.
Recently, the American Society for Biochemistry and Molecular Biology chose Drennan as the recipient of the 2023 William C. Rose Award for her outstanding contributions to biochemical research and commitment to training younger scientists. Among her additional honors are the Everett Moore Baker Memorial Award for Excellence in Undergraduate Teaching, the Harold E. Edgerton Faculty Achievement Award, the Dean’s Educational and Student Advising Award, a Committed to Caring Award, and a Presidential Early Career Award for Scientists and Engineers (PECASE). She has also been named an MIT MacVicar Fellow, a AAAS fellow, an ASBMB fellow, an Alfred P Sloan Fellow, and a Searle Scholar, and she is a member of the American Academy of Arts and Sciences.
Dina Katabi
Dina Katabi is the Thuan and Nicole Pham Professor of Electrical Engineering and Computer Science (EECS), director of the MIT Center for Wireless Networks and Mobile Computing, and a principal investigator at both the Computer Science and Artificial Intelligence Laboratory (CSAIL) and the Abdul Latif Jameel Clinic for Machine Learning in Health (Jameel Clinic), and a co-founder of Emerald Innovations. At CSAIL, she conducts mobile computing, machine learning, and computer vision research while leading the NETMIT group. Katabi is known for her contributions to wireless data transmission, developing wireless devices that assist with digital health using AI and radio signals. These works include an in-home wireless device that continuously monitors the gait speed of patients with Parkinson’s to better track the progression of the disease, an AI model that detects Parkinson’s from individuals’ breathing patterns, and BodyCompass, a radio-frequency-based wireless device that captures sleep data without using cameras or body sensors.
Katabi received a bachelor’s of science from the University of Damascus and continued her studies at MIT, where she earned a master’s of science and a PhD in computer science. She joined EECS faculty in 2003.
She is a member of the American Academy of Arts and Sciences, the National Academy of Engineering, and the National Academy of Sciences, having received the 2013 MacArthur “genius grant” Fellowship as well as the Association for Computing Machinery Prize in Computing in 2018. Additionally, Katabi has earned the ACM Grace Murray Hopper Award, two Test of Time Awards from the ACM’s Special Interest Group on Data Communications, and a Sloan Research Fellowship.
Gregory Stephanopoulos
Gregory Stephanopoulos is the W. H. Dow Professor of Chemical Engineering and Biotechnology. His work focuses on biotechnology, specifically metabolic and biochemical engineering. His research group conducts research on various projects aiming at the development of biological production routes to chemical products and biofuels. The group is also investigating cancer as metabolic disease. He is renowned for his work in reprogramming the gene transcription network of particular bacteria in order to improve their efficiency in converting renewable raw material into valuable chemical products.
Stephanopoulos graduated from the National Technical University of Athens in 1973 with the a bachelor’s degree in chemical engineering. In 1975, he obtained his master’s degree from the University of Florida and, three years later, his PhD from the University of Minnesota. His professional career started in 1978 as assistant professor at Caltech, where he was promoted in 1984 to the rank of associate professor with tenure. In 1985, Stephanopoulos moved to MIT as a professor of chemical engineering. He was Bayer Professor between 2000 and 2006, when he was appointed to the W. H. Dow Professorship of Chemical Engineering and Biotechnology. From 1990 to 1997 he served as associate director of the Biotechnology Process Engineering Center (BPEC) at MIT. In 2016, he served as president of the American Institute of Chemical Engineers (AIChE).
Stephanopoulos has received many honors, including the 2019 Gaden Award for Biotechnology and Bioengineering, the 2017 Novozymes Award for Excellence in Biochemical and Chemical Engineering, and the 2016 Eric and Sheila Samson Prime Minister’s Prize for Innovation in Alternative Fuels. In 2010, he received the George Washington Carver Award for Innovation in Industrial Biotechnology and the ACS E. V. Murphree Award. From AIChE, he has received the R.H. Wilhelm Award (2001), the Founders Award (2007) and the William Walker Award (2014). In 2011, he received the Eni Prize in Renewable and Non-Conventional Energy, and in 2013 the John Fritz Medal from the American Association of Engineering Societies. Stephanopoulos is a member of the National Academy of Engineering and a corresponding member of the Academy of Athens.

Two faculty members from the MIT Department of Biology have been selected by the Howard Hughes Medical Institute (HHMI) for the inaugural cohort of HHMI Freeman Hrabowski Scholars.
Seychelle Vos, the Robert A. Swanson Career Development Professor of Life Sciences, and Hernandez Moura Silva, an assistant professor of biology and core member of the Ragon Institute of MGH, MIT and Harvard, are among 31 early-career faculty selected for their potential to become leaders in their research fields and to create diverse and inclusive lab environments in which everyone can thrive, according to a press release.
Freeman Hrabowski Scholars are appointed to a five-year term, renewable for a second five-year term after a successful progress evaluation. Each scholar will receive up to $8.6 million over 10 years, including full salary, benefits, a research budget, and scientific equipment. In addition, they will participate in professional development to advance their leadership and mentorship skills.
The Freeman Hrabowski Scholars Program represents a key component of HHMI’s diversity, equity, and inclusion goals. Over the next 20 years, HHMI expects to hire and support up to 150 Freeman Hrabowski Scholars — appointing roughly 30 scholars every other year for the next 10 years. The institute has committed up to $1.5 billion for the Freeman Hrabowski Scholars to be selected over the next decade. The program was named for Freeman A. Hrabowski III, president emeritus of the University of Maryland at Baltimore County, who played a major role in increasing the number of scientists, engineers, and physicians from backgrounds underrepresented in science in the United States.
Seychelle Vos
Seychelle Vos studies how DNA organization impacts gene expression at the atomic level, using cryogenic electron microscopy (cryo-EM), X-ray crystallography, biochemistry, and genetics. Human cells contain about 2 meters of DNA, which is packed so tightly that its entirety is contained within the nucleus, which is only a few microns across. Although DNA needs to be compacted, it also needs to be accessible to, and readable by, the cell’s molecular machinery.
Vos received a BS in genetics from the University of Georgia in 2008 and a PhD from University of California at Berkeley in 2013. During her postdoctoral research at the Max Planck Institute for Biophysical Chemistry in Germany, she determined how the molecular machine responsible for gene expression is regulated near gene promoters.
Vos joined MIT as an assistant professor of biology in fall 2019.
“I am very humbled and honored to have been named a HHMI Freeman Hrabowski Scholar,” Vos says. “It would not have been possible without the hard work of my lab and the help of my colleagues. It provides us with the support to achieve our ambitious research goals.”
Hernandez Moura Silva
Hernandez Moura Silva studies the role of immune cells in the maintenance and normal function of our bodies and tissues, beyond their role in battling infection. Specifically, he looks at a specific type of immune cell called a macrophage and its role in the proper function of white adipose tissue — our fat. White adipose tissue in a healthy state is highly populated by macrophages, including very abundant ones known as “vasculature-associated adipose tissue macrophages,” which are located around the blood vessels. When the activity of these adipose macrophages is disrupted, there are changes in the proper function of the white adipose tissue, which may ultimately link to disease. By understanding macrophage function in healthy tissues, Hernandez hopes to learn how to restore tissue homeostasis in disease.
Hernandez Moura Silva received a BS in biology in 2005 and an MSc in molecular biology in 2008 from the University of Brazil. He received his PhD in 2011 from the University of São Paulo Heart Institute. Silva pursued his postdoctoral work as the Bernard Levine Postdoctoral Fellow in immunology and immuno-metabolism at the New York University School of Medicine Skirball Institute of Biomolecular Medicine.
He joined MIT as an assistant professor of biology in 2022. He is also a core member of the Ragon Institute.
“For an immigrant coming from an underrepresented group, it’s a huge privilege to be granted this opportunity from HHMI that will empower me and my lab to shape the next generation of scientists and provide an environment where people can feel welcome and encouraged to do the science that they love and be successful,” Silva says. “It also aligns with MIT’s commitment to increase diversity and opportunity across the Institute and to become a place where all people can thrive.”

Billions of times a day, every day of our lives, cells receive signals to initiate the process of cell death. This strategic cell death, also called apoptosis, is one of the tools multicellular organisms use to maintain tissues and regulate immune responses: damaged, old, or superfluous cells are given the green light to, as it were, turn out the lights for the last time.
Programmed cell death is both extremely powerful and extremely regulated: for example, the careful culling of cells between our digits during embryonic development reveals fingers and toes. When programmed cell death goes awry, however, it can have serious consequences. Cells left unchecked can divide unstoppably and aggressively, leading to cancer. Dysregulated apoptotic pathways have also been implicated in neurodegenerative diseases like Alzheimer’s, where unrestrained cell death may play a part in the severity of the disease.
MIT Professor H. Robert Horvitz ‘68 shared a Nobel prize in 2002 for his foundational research on the genetics of programmed cell death and organ development in the nematode, a microscopic roundworm. Horvitz discovered that ced-9, a key gene in programmed cell death in nematodes, was similar in structure and function to the human gene bcl-2.
Targeting members of the BCL-2 protein family has already shown promise in the fight against cancer. For example, approved by the FDA in 2016, the oral drug Venetoclax is a BCL-2 inhibitor used to treat certain types of leukemia.
In a study published online Jan. 26 in Structure, Fiona Aguilar PhD ‘22 (Keating lab) and collaborators focused on a member of the BCL-2 protein family called BAK. When it is active, BAK promotes mitochondrial outer membrane disruption, leading to cell death, and is therefore referred to as a pro-apoptotic protein. But precisely how BAK becomes activated – or inhibited – is unknown.
“A greater understanding of BAK activation is interesting both from a fundamental biochemical and biophysical perspective as well as from the more translational one of BAK as a potential therapeutic target,” says lead author Fiona Aguilar.
BAK exists in two different forms: an inactive monomer and an active oligomer. A few activators of BAK (BIM, truncated BID, and PUMA) have already been identified and these proteins bind directly to BAK, leading to the model that binding of activators trigger changes in protein shape that allow BAK to transition from the inactive to active forms. To further explore this idea, Aguilar identified and characterized a number of other peptides that bind to and regulate BAK. To identify new peptide binders, the team used cell-surface display screening and computational protein design methods, including techniques developed by Keating lab alum Gevorg Grigoryan– dTERMen and TERMify – that use protein structural data to generate new protein sequences likely to bind a protein of interest.
In total, Aguilar et al. discovered 10 diverse new peptide binders of BAK that regulate its function.
Interestingly, some of the BAK-binding peptides inhibited activation rather than promoting it. Aguilar et al. found that inhibitors and activators of BAK shared many characteristics including structure as well as binding affinity and kinetics – the strength and rate that binders associate with and dissociate from BAK.
Newly identified activators had sequences both dissimilar from one another and from the previously known BAK activators BIM, truncated BID, and PUMA. The similarity of the sequence was not necessarily a good indicator of activation or inhibition. For example, an inhibitor and an activator differed by just two amino acids.
Aguilar and colleagues solved the crystal structures of two inhibitor-BAK complexes and one activator-BAK complex and found that the activator interacted with BAK with similar geometry as the two inhibitors. Also, the two inhibitors have only about 40% sequence identity, but bind very similarly to BAK.
Amy Keating, the senior author on the study, says “Fiona was tireless in identifying new peptides, testing their interactions with BAK, determining their functions, and solving structures to look for differences between activators and inhibitors. We were surprised that peptides with such different behaviors shared such common interaction properties.”
Although the puzzle is not yet solved, Aguilar believes the “transition state” between inactive and active forms of BAK is key.
“We think of activators as peptides that preferentially bind to the BAK transition state, whereas inhibitors are those that preferentially bind to the monomeric state,” Aguilar says. “Overall, we should be thinking more about the transition state, what steps are necessary to reach the transition state, and how to target the transition state.”
This study also added two sequences in the human proteome – BNIP5 and PXT1 – to the repertoire of known BAK binders. Not much is known about these sequences, Aguilar says, but the fact that they activate BAK could indicate that they may play a role in apoptotic pathways that have not yet been determined.
“The finding is something that people in the field are pretty excited about,” Aguilar says.
Ultimately, work remains to establish what characteristics of the binders determine their function, and how binding to BAK triggers the conformational changes that activate or inhibit this complex protein.
“It’s still unclear what it is about these sequences that trigger the allosteric network leading to BAK activation, but at least for now we can rule out the hypothesis that binding mode, affinity, and kinetics fully determine how this occurs,” Aguilar says.
Aguilar suggests that it will be interesting also to explore how these peptides interact with BAX, another pro-apoptotic protein in the BCL-2 family that is both structurally and functionally similar to BAK.
Fiona Aguilar is lead author and Amy Keating is senior author; Bob Grant and graduate students Sebastian Swanson, Dia Ghose, and Bonnie Su contributed. Collaborators Stacey Yu and Kristopher Sarosiek, from the Harvard T.H. Chan School of Public Health, helped with cell-based experiments. The research was funded by a National Institute of General Medical Sciences award, the MIT School of Science Fellowship in Cancer Research award, the John W. Jarve (1978) Seed Fund for Science Innovation (MIT) award, an award from the National Cancer Institute, a National Institute of Diabetes and Digestive and Kidney Diseases award, and Alex’s Lemonade Stand Foundation for Childhood Cancers award.

How does animal behavior emerge from networks of connected neurons? How are these incredible nervous systems and behaviors actually generated by evolution? Are there principles shared by all nervous systems or is evolution constantly innovating? What did the first nervous system look like that gave rise to the incredible diversity of life that we see around us?
Combining the study of animal behavior with studies of nervous system form, function, and evolution, Brady Weissbourd, a new faculty member in the Department of Biology and investigator in The Picower Institute for Learning and Memory, uses the tiny, transparent jellyfish Clytia hemisphaerica, a new neuroscience model.
Q: In your work, you developed a new model organism for neuroscience research, the transparent jellyfish Clytia hemisphaerica. How do these jellyfish answer questions about neuroscience, the nervous system, and evolution in ways that other models cannot?
A: First, I believe in the importance of more broadly understanding the natural world and diversifying the organisms that we deeply study. One reason is to find experimentally tractable organisms to identify generalizable biological principles – for example, we understand the basis of how neurons “fire” from studies of the squid giant axon. Another reason is that transformative breakthroughs have come from identifying evolutionary innovations that already exist in nature – for example, green fluorescent protein (GFP, from jellyfish) or CRISPR (from bacteria). In both ways, this jellyfish is a valuable complement to existing models.
I have always been interested in the intersection of two types of problems: how nervous systems generate our behaviors; and how these incredible systems were actually created by evolution.
On the systems neuroscience side, ever since working on the serotonin system during my PhD I have been fascinated by the problem of how animals control all of their behaviors simultaneously in a flexible and context-dependent manner, and how behavioral choices depend not just on incoming stimuli but on how those stimuli interact with constantly changing states of the nervous system and body. These are extremely complex and difficult problems, with the particular challenge of interactions across scales, from chemical signaling and dynamic cell biology to neural networks and behavior.
To address these questions, I wanted to move into a model organism with exceptional experimental tractability.
There have been exciting breakthroughs in imaging techniques for neuroscience, including these incredible ways in which we can actually watch and manipulate neuronal activity in a living animal. So, the first thing I wanted was a small and transparent organism that would allow for this kind of optical approach. These jellyfish are a few millimeters in diameter and perfectly transparent, with interesting behaviors but relatively compact nervous systems. They have thousands of neurons where we have billions, which also puts them at a nice intermediate complexity compared to other transparent models that are widely used – for example, C. elegans have 302 neurons and larval zebrafish have something like 100,000 in the brain alone. These features will allow us to look at the activity of the whole nervous system in behaving animals to try to understand how that activity gives rise to behaviors and how that activity itself arises from networks of neurons.
On the evolution side of our work, we are interested in the origins of nervous systems, what the first nervous systems looked like, and broadly what the options are for how nervous systems are organized and functioning: to what extent there are principles versus interesting and potentially useful innovations, and if there are principles, whether those are optimal or somehow constrained by evolution. Our last common ancestor with jellyfish and their relatives (the cnidarians) was something similar to the first nervous system, so by comparing what we find in cnidarians with work in other models we can make inferences about the origins and early evolution of nervous systems. As we further explore these highly divergent animals, we are also finding exciting evolutionary innovations: specifically, they have incredible capabilities for regenerating their nervous systems. In the future, it will be exciting to better understand how these neural networks are organized to allow for such robustness.
Q: What work is required to develop a new organism as a model, and why did you choose this particular species of jellyfish?
A: If you’re choosing a new animal model, it’s not just about whether it has the right features for the questions you want to ask, but also whether it technically lets you do the right experiments. The model we’re using was first developed by a research group in France, who spent many years doing the really hard work of figuring out how to culture the whole life cycle in the lab, injecting eggs, and developing other key resources. For me, the big question was whether we’d be able to use the genetic tools that I was describing earlier for looking at neural activity. Working closely with collaborators in France, our first step was figuring out how to insert things into the jellyfish genome. If we couldn’t figure that out, I was going to switch back to working with mice. It took us about two years of troubleshooting, but now we can routinely generate genetically modified jellyfish in the lab.
Switching to a new animal model is tough – I have a mouse neuroscience background and joined a postdoc lab that used mice and flies; I was the only person working with jellyfish but had no experience. For example, building an aquaculture system and figuring out how to keep jellyfish healthy is not trivial, particularly now that we’re trying to do genetics. One of my goals is now to optimize and simplify this whole process so that when other labs want to start working with jellyfish we have a simple aquaculture platform to get them started, even if they have no experience.
In addition to the fact that these things are tiny and transparent, the main reason that we chose this particular species is because it has an amazing life cycle that makes it an exciting laboratory animal.
They have separate sexes that spawn daily with the fertilized eggs developing into larvae that then metamorphose into polyps. We grow these polyps on microscope slides, where they form colonies that are thought to be immortal. These colonies are then constantly releasing jellyfish, which are all genetically identical “clones” that can be used for experiments. That means that once you create a genetically modified strain, like a transgenic line or a knockout, you can keep it forever as a polyp colony – and since the animals are so small, we can culture them in large numbers in the lab.
There’s still a huge amount of foundational work to do, like characterizing their behavioral repertoire and nervous system organization. It’s shocking how little we know about the basics of jellyfish biology – particularly considering that they kill more people per year than sharks and stingrays combined – and the more we look into it the more questions there are.
Q: What drew you to a faculty position at MIT?
A: I wanted to be in a department that does fundamental research, is enthusiastic about basic science, is open-minded, and is very diverse in what people work on and think about. My goal is also to be able to ultimately link mechanisms at the molecular and cellular level to organismal behavior, which is something that MIT Biology is particularly strong at doing. It’s been an exciting first few months! MIT Biology is such an amazing place to do science and it’s been wonderful how enthusiastic and supportive everyone in the department has been.
I was additionally drawn to MIT by the broader community and have already found it so easy to start collaborations with people in neuroscience, engineering, and math. I’m also thrilled to have recently become a member of The Picower Institute for Learning and Memory, which further enables these collaborations in a way that I believe will be transformational for the work in my lab.
It’s a new lab. It’s a new organism. There isn’t a huge, well-established field that is taking these approaches. There’s so much we don’t know, and so much that we have to establish from scratch. My goal is for my lab to have a sense of adventure and fun, and I’m really excited to be doing that here in MIT Biology.