Drug that targets a key cancer protein could combat leukemia and other types of cancer.
Anne Trafton | MIT News Office
January 15, 2018
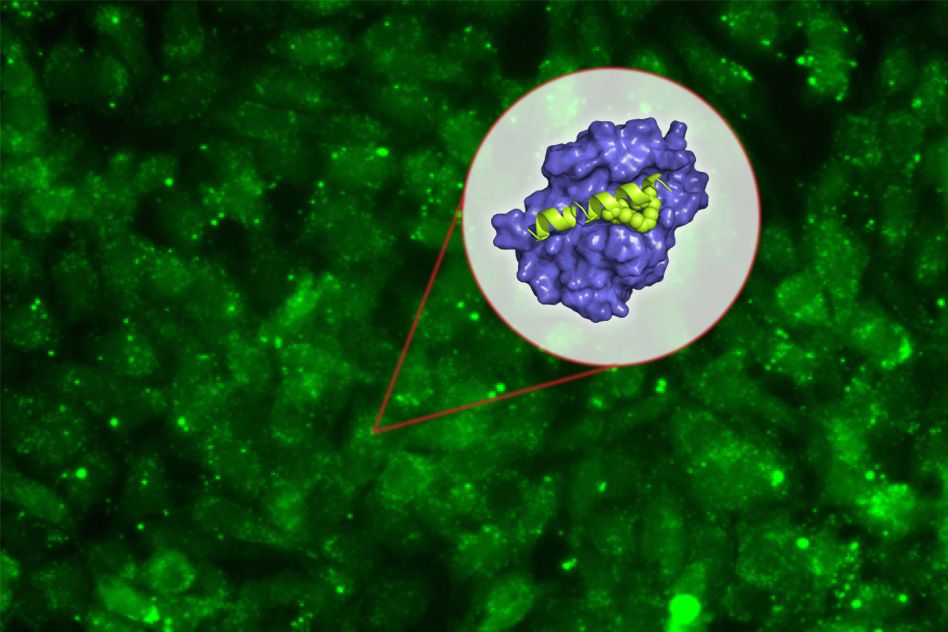
MIT biologists have designed a new peptide that can disrupt a key protein that many types of cancers, including some forms of lymphoma, leukemia, and breast cancer, need to survive.
The new peptide targets a protein called Mcl-1, which helps cancer cells avoid the cellular suicide that is usually induced by DNA damage. By blocking Mcl-1, the peptide can force cancer cells to undergo programmed cell death.
“Some cancer cells are very dependent on Mcl-1, which is the last line of defense keeping the cell from dying. It’s a very attractive target,” says Amy Keating, an MIT professor of biology and one of the senior authors of the study.
Peptides, or small protein fragments, are often too unstable to use as drugs, but in this study, the researchers also developed a way to stabilize the molecules and help them get into target cells.
Loren Walensky, a professor of pediatrics at Harvard Medical School and a physician at Dana-Farber Cancer Institute, is also a senior author of the study, which appears in the Proceedings of the National Academy of Sciences the week of Jan. 15. Researchers in the lab of Anthony Letai, an associate professor of medicine at Harvard Medical School and Dana-Farber, were also involved in the study, and the paper’s lead author is MIT postdoc Raheleh Rezaei Araghi.
A promising target
Mcl-1 belongs to a family of five proteins that play roles in controlling programmed cell death, or apoptosis. Each of these proteins has been found to be overactive in different types of cancer. These proteins form what is called an “apoptotic blockade,” meaning that cells cannot undergo apoptosis, even when they experience DNA damage that would normally trigger cell death. This allows cancer cells to survive and proliferate unchecked, and appears to be an important way that cells become resistant to chemotherapy drugs that damage DNA.
“Cancer cells have many strategies to stay alive, and Mcl-1 is an important factor for a lot of acute myeloid leukemias and lymphomas and some solid tissue cancers like breast cancers. Expression of Mcl-1 is upregulated in many cancers, and it was seen to be upregulated as a resistance factor to chemotherapies,” Keating says.
Many pharmaceutical companies have tried to develop drugs that target Mcl-1, but this has been difficult because the interaction between Mcl-1 and its target protein occurs in a long stretch of 20 to 25 amino acids, which is difficult to block with the small molecules typically used as drugs.
Peptide drugs, on the other hand, can be designed to bind tightly with Mcl-1, preventing it from interacting with its natural binding partner in the cell. Keating’s lab spent many years designing peptides that would bind to the section of Mcl-1 involved in this interaction — but not to other members of the protein family.
Once they came up with some promising candidates, they encountered another obstacle, which is the difficulty of getting peptides to enter cells.
“We were exploring ways of developing peptides that bind selectively, and we were very successful at that, but then we confronted the problem that our short, 23-residue peptides are not promising therapeutic candidates primarily because they cannot get into cells,” Keating says.
To try to overcome this, she teamed up with Walensky’s lab, which had previously shown that “stapling” these small peptides can make them more stable and help them get into cells. These staples, which consist of hydrocarbons that form crosslinks within the peptides, can induce normally floppy proteins to assume a more stable helical structure.
Keating and colleagues created about 40 variants of their Mcl-1-blocking peptides, with staples in different positions. By testing all of these, they identified one location in the peptide where putting a staple not only improves the molecule’s stability and helps it get into cells, but also makes it bind even more tightly to Mcl-1.
“The original goal of the staple was to get the peptide into the cell, but it turns out the staple can also enhance the binding and enhance the specificity,” Keating says. “We weren’t expecting that.”
Killing cancer cells
The researchers tested their top two Mcl-1 inhibitors in cancer cells that are dependent on Mcl-1 for survival. They found that the inhibitors were able to kill these cancer cells on their own, without any additional drugs. They also found that the Mcl-1 inhibitors were very selective and did not kill cells that rely on other members of the protein family.
Keating says that more testing is needed to determine how effective the drugs might be in combating specific cancers, whether the drugs would be most effective in combination with others or on their own, and whether they should be used as first-line drugs or when cancers become resistant to other drugs.
“Our goal has been to do enough proof-of-principle that people will accept that stapled peptides can get into cells and act on important targets. The question now is whether there might be any animal studies done with our peptide that would provide further validation,” she says.
Joshua Kritzer, an associate professor of chemistry at Tufts University, says the study offers evidence that the stapled peptide approach is worth pursuing and could lead to new drugs that interfere with specific protein interactions.
“There have been a lot of biologists and biochemists studying essential interactions of proteins, with the justification that with more understanding of them, we would be able to develop drugs that inhibit them. This work now shows a direct line from biochemical and biophysical understanding of protein interactions to an inhibitor,” says Kritzer, who was not involved in the research.
Keating’s lab is also designing peptides that could interfere with other relatives of Mcl-1, including one called Bfl-1, which has been less studied than the other members of the family but is also involved in blocking apoptosis.
The research was funded by the Koch Institute Dana-Farber Bridge Project and the National Institutes of Health.