July 7, 2019
Unusual Labmates is a series that explores some of the more unusual models used for research at Whitehead Institute. From rare plants to luminescent beetles to regenerative starfish and worms, these organisms and their unusual traits provide insights into the underlying biology and incredible diversity of living things.
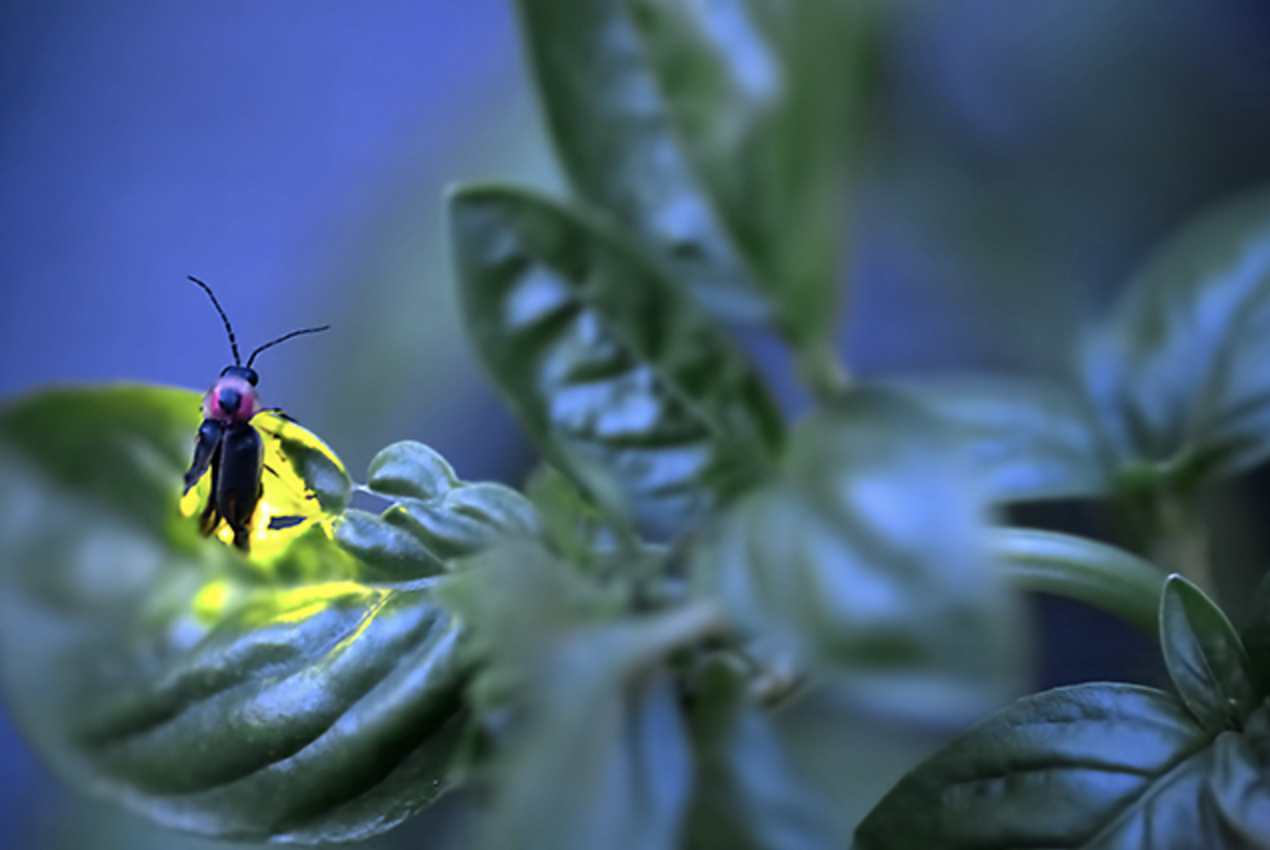
Massachusetts Institute of Technology (MIT) graduate student Tim Fallon is standing in a field in New Jersey, holding a net and waiting for the last glimmers of sunlight to disappear. As the trees surrounding the field fade into shadow, Fallon watches the ground intently. The air is still and then, hovering above the grass, a small point of light appears. It floats through the air in a concave upwards arc and winks out. Soon, more lights follow suit. These are fireflies, unassuming beetles by day, but at night they put on a dazzling luminescent display in the hope of attracting a mate. Fallon sweeps his net through the air, capturing some of them. He has traveled to New Jersey with other members of Whitehead Institute Member and associate professor of biology at MIT Jing-Ke Weng’s lab to collect specimens of Photinus pyralis, the big dipper firefly, in order to sequence a firefly genome for the very first time.
Fireflies have been around for more than one hundred million years, and in that time have diverged into more than 2,000 species and spread to every continent except Antarctica. The beetles (despite their name, fireflies are actually beetles) are widely known, and often beloved, for their enchanting courtship rituals, but they have also piqued the interest of scientists, who have harnessed the gene behind their light emitting capability for use in research. However, for all of fireflies’ appeal, they are a difficult animal to work with in the lab, and much of their biology remains shrouded in mystery. In the hopes of improving that situation, Weng, Fallon, and collaborators—including Sarah Lower, assistant professor of biology at Bucknell University, and Yuichi Oba, professor at Chubu University in Japan—have been investigating fireflies, primarily by sequencing and analyzing their genomes. This research is providing insights into the evolution of fireflies’ light-producing ability, the biomolecular pathways the fireflies use to luminesce, and could perhaps even inform how the use of firefly-based luminescence in research can be improved.
The chemistry of light
Fireflies produce light using two main ingredients: an enzyme called luciferase and the small molecule luciferin. Luciferase facilitates a chemical reaction that oxidizes the luciferin, and one product of the reaction is light. In the 1980s, researchers recreated the process in plants and plant cells by cloning the firefly gene responsible for the luciferase enzyme from big dipper fireflies and then inserting it into the genomes of their specimens in the lab.[1] When they injected luciferin into the specimens, they began to glow – just like fireflies. Researchers now use this approach in both plant and animal models to track various aspects of biology. They can link the luminescence to a trait or process, and then measure the level of light emitted. For example, researchers can fuse the luciferase gene to a gene of interest such that the two genes will be expressed as one. Then they introduce luciferin to the system and measure the light output using very sensitive equipment that can sense minute changes. The more light that is emitted, the higher the activity level of both luciferase and the gene of interest. This is a widely used assay that, Fallon says, every biologist uses or at least learns about during their training. Luciferase-luciferin has many applications, and along with tracking gene expression it has also been used to track cancer metastasis, monitor medical treatment efficacy, and check for microbial contamination—including on space vehicles such as the Mars Curiosity Rover.[2] The extreme sensitivity of luciferin-luciferase tests makes them an attractive choice in many experiments.
In spite of the widespread use of firefly luminescence, a lot about it still isn’t known, especially when it comes to luciferin. While the gene encoding luciferase has been identified, luciferin is thought to be created through a process involving multiple genes, and the complete set of those genes is unknown, as are the steps involved in the production of luciferin: the intermediate molecules produced and how they are modified to reach the final product. Although Weng’s lab generally studies plants and focuses on understanding how plants evolved biochemical pathways to produce unique small molecules with traits of interest, particularly those with medicinal value, when Fallon joined the lab, he convinced Weng that investigating the unknowns of the small molecule luciferin was a similar and suitable project.
Fireflies in the wild
When Fallon joined the Weng lab, there were not a lot of research tools available for investigating fireflies, starting with the lack of a sequenced genome. Only a handful of firefly genes had even ever been identified. Weng and his collaborators crowdfunded the money to sequence the first firefly genome, and then received funding to sequence a second firefly species and a bioluminescent click beetle, providing a wealth of new data to explore.
The lack of tools for firefly research was due in part to how difficult it is to rear fireflies in the lab, which makes them tricky animals to study. Fireflies are very sensitive to changes in their environments, and in the lab it’s difficult to mimic the right vegetation, climate, seasonal shifts, and other factors that the beetles rely on to time their metamorphoses and thrive.
Unpredictable environmental changes are becoming a common challenge for fireflies beyond the lab as well, due to the impact of humans on their habitats. Fireflies’ sensitivity to these changes may be causing their disappearance. Anecdotally, many people have fond memories of seeing fireflies every summer when they were younger and can attest to the absence of fireflies in those same places now. A large citizen science research project called Firefly Watch is currently underway to figure out the extent to which firefly populations are declining across the United States (U.S.).[3] And in a case indicative of a larger problem, conservation groups recently submitted an urgent petition to recognize the Bethany Beach firefly, whose key habitat in Delaware is at risk due to human development, as the first endangered firefly species in the U.S.[4]
A significant manmade disruption to the fireflies’ habitats is light pollution. Fireflies find each other by signaling with light, but their lanterns are no match for electricity; a firefly trying to outshine a streetlamp might as well be facing off with the sun. For species that evolved faint glows to flash in the dead of night, in the dark of forests, finding a place where their light can be seen is getting harder and harder. Big dipper fireflies have fared better than many others species. They’re large, with bright lanterns, and they tend to flash at twilight, so they are used to competing with some ambient light. Unsurprisingly, big dipper fireflies remain abundant in the wild, including in and near cities. They can be spotted all over the eastern and midwestern U.S., where they are easily identifiable thanks to the distinctive J-shape—resembling the curve of a dipper—that they make as they flash.
Big dipper fireflies have been important in biology research—it is from them that the firefly luciferase gene was first cloned—and so they were the first species that Fallon and his collaborators chose for their genome sequencing project. However, big dipper fireflies fare no better than other firefly species in a lab setting. No one has ever successfully reared big dipper fireflies through a full lifecycle (from egg to egg) in the lab, Fallon says. So, in spite of the ease of collecting big dipper fireflies in the eastern U.S., in order to get a population of fireflies going in the lab, Fallon had to look elsewhere: to Japan.
In Japan, fireflies are beloved. Watching them is a popular summer pastime, and they are celebrated in myths, songs, and other media. The two dominant species are both aquatic in their larval stages: the heike-botaru (Aquatica lateralis), which mostly lives in flooded rice paddy fields, and the larger and brighter genji-botaru (Luciola cruciata), which lives in streams and rivers. Both species need clean bodies of water to survive, and so their numbers have diminished in cities—but unlike in America, Japan has not allowed fireflies to fade away quietly into the night. As fireflies have grown scarcer, breeding centers have begun rearing the insects in large numbers, using big facilities that can recreate the fireflies’ natural habitat on a scale not possible in a U.S. lab. Fireflies are sometimes bred for conservation purposes, such as an effort to bolster the heike population in the water by the Imperial Palace in Tokyo. The fireflies are also bred for spectacle, released during summer festivals and firefly-watching events in the city to recreate the lost experience of glow-filled nights. The Fussa firefly festival has drawn large crowds for more than fifty years.
Rearing insects is also a relatively common pastime in Japan. When Fallon was looking for a type of firefly that would be a good addition to the genome sequencing project and could survive in lab conditions, he learned of the heike, which are kept in captivity and used in research in Japan. Although heike are still very sensitive to small changes in their environments, they have been demonstrated to survive for multiple generations indoors, unlike the big dipper firefly. The larvae are aquatic, and maintaining a controlled aquarium is also easier than a terrarium, Fallon says. Fallon received his fireflies from a collaborator in Japan, firefly expert Dr. Yuichi Oba, a professor at Chubu University. Oba works with Haruyoshi Ikeya, a high school teacher in Yokohama, Japan who cracked the code of rearing heike fireflies indoors several decades ago; the population from which Fallon received his specimens has been lab-bred since its original capture in 1990. Fallon received tips from Oba on how to successfully rear the fireflies—though not all of these could be followed, due to stricter regulations for how to keep the fireflies contained in the U.S., where the United States Department of Agriculture considers them a potential plant pest.
Fallon rears the fireflies in a small room separate from the main space of the Weng lab. The room is tightly packed with supplies and various aquariums: Fallon must move the firefly specimens between environments as they go through their lifecycle. There is tinfoil over the windows so when Fallon switches the lights off the room becomes a darkroom, where he can observe the fireflies flashing.
Maintaining a lab population serves a number of purposes. First of all, collecting fireflies is time-consuming, limited by season, and in the case of foreign species, involves further transportation and regulatory roadblocks. When collecting specimens in the wild, it’s easy to find adults, but harder to get access to every stage of the lifecycle, especially eggs and pupae, the way you can in the lab. Having a lab population also ensures that you are working with one species, whereas when collecting in the wild it can be easy to get a mixed-up batch—one field may contain a dozen similar-looking species—and molecular biology research requires species-specific material. Overall, having a lab population means access to live specimens whenever you need them: whenever the researchers have a new research question to answer or a new experiment to run. However, maintaining this beneficial resource is not a simple feat.
Rearing fireflies in captivity is difficult in part because each stage of their life cycle has different requirements. Fireflies hatch from eggs, which the heike lay in moss near water. Heike larvae live in water, where they spend most of their lifespan, usually around one year, though in the lab this stage can be as short as six months. During this time, the larvae typically go through five or six instars, or stages of growth between molting. Insects must molt in order to grow, as their size is restricted by their exoskeletons. The heike’s first instar is only a few millimeters long, while the final instar may grow to be closer to two centimeters.
For most of their time as larvae, the specimens live in shoebox sized aquariums inside a metal cabinet. Fallon feeds them bladder snails and waits for them to grow. When the larvae are in their later instars, around the right size and age to pupate, Fallon moves them into another aquarium with a mock riverbank inside, with a soil mixture devised by Haruyoshi Ikeya. The riverbank is necessary because the firefly larvae move to land to pupate. First, they begin to exhibit what is called climbing or landing behavior, during which they will flash—fireflies are capable of emitting light at all stages of their lifecycle. The flashes may be a signal to help coordinate pupation. If a larva doesn’t synchronize with the others when it is ready to pupate, there won’t be any mates around when it hatches. Fireflies don’t live very long as adults, so they must find a mate quickly. In the lab, once the adults hatch, Fallon moves them into another container to prevent them from drowning in the water of the mock river, and hopes that they mate. If they are successful, then the female lays her eggs in the moss Fallon provides, starting the cycle over again.
The fireflies are not easy to keep alive in lab conditions, so the researchers have been experimenting with different conditions for rearing them, like keeping them in boxes of different sizes, changing the aeration of their water, and providing different spaces for the adults to rest when they first hatch.
“We’ve been iterating through a lot of different ways to rear them,” Fallon says. “It’s not like with fruit flies, where you could leave two alone in a room with a banana, and soon you’ve got more fruit flies than you know what to do with. The fireflies are really quite finicky. Over time we’re learning what works and what doesn’t.”
One thing that Fallon has learned from rearing fireflies is how often they glow, during every part of their life cycle. The heike lay their eggs in clumps, which are cumulatively visible to the naked eye. Big dipper firefly eggs also luminesce, but faintly enough that it’s only visible using a sensitive camera. If there’s an evolutionary advantage to having the eggs luminesce, it’s not known. The larvae’s ability to glow is typically believed to be, at least in part, an aposematic signal: a warning to potential predators that the larvae are toxic so the predators won’t try to eat them. Other species use bright colors for the same purpose—think of brilliantly colored poison dart frogs or the popular mnemonic “red touches yellow kills a fellow” used to identify venomous coral snakes—and what’s brighter than something that glows in the dark?
Fireflies’ toxicity comes from chemicals called lucibufagins. Only some firefly species produce these, though the rest may benefit by association if predators associate glowing beetles with a bitter meal. Fallon has noticed that the heike larvae will glow anytime they appear threatened or are faced with the unfamiliar, like if he jostles the box they live in. This threat response is consistent with the expectations for an aposematic signal, but the heike larvae also glow in other conditions, such as during their climbing behavior right before they pupate—and the pupae glow as well. The adults, meanwhile, can also warn off predators with their flashes but primarily luminesce to communicate and find a mate. Different firefly species have distinct flash patterns, as do males and females, allowing adults to identify a suitable partner—and allowing anyone with enough firefly knowledge to identify different species just by watching their flashes.
In every stage of life, fireflies have adapted to make the most of their light-emitting ability. Once they had the genome sequences and firefly specimens in hand, Weng and Fallon wanted to find out how.
The researchers used the firefly genomes to delve into the biomolecular pathways that fireflies use to create light. They identified a luciferin-derived molecule in fireflies that may be a storage form for luciferin, as well as the gene encoding the enzyme that converts luciferin into this molecule. They have also been looking into fireflies’ mechanism for recycling luciferin. The one significant handicap of using luciferase-luciferin in research is that, since the genes encoding luciferin are unknown, the chemical must be fed or injected into specimens manually. This limits the duration of experiments based on when the luciferin is used up, or requires disrupting specimens to reinject them. In the wild, fireflies are able to reuse their luciferin molecules after they have been oxidized to produce light; if researchers could figure out how they do this, it could potentially be applied in the lab.
In order to solve this puzzle, the researchers sequenced the genome of three beetles and compared them: the big dipper firefly, the heike firefly, and the cucubano click beetle (Ignelater luminosus), which is also capable of luminescence. The last common ancestor of heike and big dipper fireflies lived over 100 million years ago, making them good candidates for evolutionary comparison. The researchers found evidence that fireflies and click beetles evolved luminescence independently. They pinpointed where the luciferase gene was in each species’ genome, and what its neighbors were—in the fireflies, the gene was surrounded by genes involved in fatty acid metabolism, suggesting that it evolved from one of these. Meanwhile, the researchers found the click beetle’s luciferase gene in a completely different genetic neighborhood, suggesting that it evolved separately and from a different ancestral gene.
It may seem unusual for such an extraordinary trait as luminescence to have evolved multiple times, but in fact it is a trait that evolution constantly stumbles upon, Fallon says. Beetles are far from the only creatures capable of emitting light; bioluminescence has also evolved in many niches, including in species of fish, coral, jellyfish, squid, snails, fungi, and bacteria.
Fireflies’ luminescence is not a unique trait, but it’s one worth preserving. From fireflies lighting up the night sky at summer festivals and in backyards, to children chasing them through the grass trying to capture a little magic in their hands, to researchers exploring biology with the help of the ultra-sensitive luciferase gene, people benefit from sharing our world with these dazzling little beetles. With the new data coming out of labs like Weng’s, further research benefits from fireflies’ light-making machinery may be on the horizon.
Fallon has learned a lot about the difficulties of rearing fireflies as he tries to maintain a sustainable population in the lab; meanwhile conservationists are struggling to protect populations of fireflies out in the wild. Even the wild population from which Fallon’s fireflies were originally captured no longer exists. Though the species survives, that particular population’s habitat disappeared, leaving the lab-bred beetles as their only legacy. The more that researchers learn about fireflies, the better equipped we may be to protect them from the sort of environmental vulnerabilities that killed off the Weng lab fireflies’ ancestors—both for the sake of the fireflies themselves, and for own sake as spectators and researchers.
Credits
Written by Greta Friar
Video by Conor Gearin
Audio production by Conor Gearin
Cover video by Radim Schreiber / FireflyExperience.org
“The chemistry of light” title card photo by Tim Fallon
“Fireflies in the wild” title card photo by Radim Schreiber / FireflyExperience.org
“Fireflies in the lab” title card photo by Conor Gearin
“What can we learn from fireflies?” title card photo by Conor Gearin
Special thanks to Radim Schreiber, Tim Fallon and Jing-Ke Weng
Works cited:
[1] https://science.sciencemag.org/content/234/4778/856
[2] https://www.science.gov/topicpages/p/planetary+protection+protocols
[3] https://www.massaudubon.org/get-involved/citizen-science/firefly-watch
[4] https://xerces.org/2019/05/15/bethany-beach-firefly/