“Selfish” DNA helps bacteria cheat and grow in densely-packed microbial communities
Raleigh McElvery
Scientists have a term for genes that spread themselves throughout a population at any cost: “selfish” DNA. One way that these genes transmit through bacterial communities is via a type of bacterial sex called conjugation. When one bacterium makes contact with another, DNA from the host cell can be injected into a recipient cell.
Alan Grossman’s lab at the MIT Department of Biology studies a small but selfish chunk of DNA called ICEBs1. His group has identified several ways in which this so-called mobile genetic element actually benefits its host bacterium as it fights to spread. Building off this body of work, Grossman’s lab collaborated with colleagues at Tel Aviv University on a new study recently published in eLife. The international team found that ICEBs1 contains one gene in particular, which allows the host cell to continue dividing in densely-packed microbial communities. This helps the host to grow in conditions where nutrients are scarce, while also potentially helping ICEBs1 to propagate.
“Mobile genetic elements like ICEBs1 are found in the chromosomes of many different types of bacteria,” says Grossman, department head and co-senior author on the study. “Studying these elements — how they spread and how they affect their host cells — is critical for understanding the evolution of bacteria, engineering some types of bacteria to do useful things, and possibly preventing the deleterious effects caused by harmful bacteria.”
Like many DNA segments on the move, ICEBs1 includes genes that encode the molecular machinery required to transfer itself from one cell to the next. But mobile genetic elements can also contain “cargo” genes that bestow the host bacterium with new traits, such as antibiotic resistance. However, in many cases, the properties a cargo gene will endow are hard to predict.
“The host cell can get a lot of new genes in a hurry through mobile genetic elements like ICEBs1, and there’s a lot we still don’t know about the types of phenotypes cargo genes confer,” says the study’s first author, Joshua Jones PhD ’20. “The array of possible traits is probably a lot more diverse than we currently appreciate.”
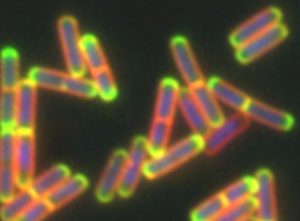
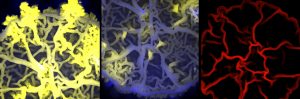
To investigate the changes that ICEBs1 triggers in the host cell, Jones and colleagues examined large microbial communities called biofilms. These form when many bacteria aggregate on a surface and secrete a slimy “glue” made of sugar, proteins, and DNA that encases the population. Common examples of biofilms include dental plaque, the sludge that coats the inside of pipes, or the deleterious infections that form on surgical implants in patients’ bodies.
Because there are so many bacteria in close contact, biofilms are hot spots for exchanging mobile genetic elements like ICEBs1. However, secreting the materials needed to produce the slimy glue can rapidly deplete resources. As a result, bacteria in a biofilm do not always have the capacity to grow, divide, and potentially spread ICEBs1. Instead, certain types of rod-shaped bacteria begin to produce spores that are analogous to plant seeds. This process, called sporulation, enables these bacteria to become dormant and survive extreme conditions.
Jones found that Bacillus subtilis bacteria containing ICEBs1 were delayed in contributing to the biofilm glue, and also delayed in producing dormant spores. As a result, these bacteria could continue dividing for longer than bacteria without ICEBs1 — increasing the number of bacteria with ICEBs1 and the likelihood that ICEBs1 would spread. The researchers were able to pinpoint one ICEBs1 cargo gene in particular, called Development Inhibitor (devI), that triggered this delay in both biofilm development and sporulation.
“In a way, the cells with ICEBs1 are ‘cheating’ by delaying sporulation and not contributing to the greater good of the biofilm community,” Jones says. But, he explains, they can get away with it because the devI pathway only initiates when ICEBs1-containing cells are the minority in a microbial population. In order to spread as widely as possible, it’s best for ICEBs1 to transfer to new cells that don’t already contain existing copies. Furthermore, accumulating duplicate copies can have detrimental effects on ICEBs1 itself.
“It’s a very clever system for assessing the situation around the cell, and deciding whether it’s worthwhile for ICEBs1 to attempt to transfer,” Jones adds.
Next, the Grossman lab plans to determine precisely how devI exerts its effects on biofilm formation and sporulation. They suspect that other ICEBs1-like elements may also use genes analogous to devI to execute similar propagation strategies. Probing such “cheating” tactics orchestrated by selfish genes will help scientists better understand microbial evolution and, eventually, perhaps even inspire drugs to disrupt harmful biofilms, like those that form around surgical implants.
Top image: Three biofilms. Credit: llana Grinberg.
Citation:
“A mobile genetic element increases bacterial host fitness by manipulating development”
eLife, online Mar 3, 2021, DOI: 10.7554/eLife.65924
Joshua M Jones, Ilana Grinberg, Avigdor Eldar, and Alan D Grossman