Forging a new understanding of metal-containing proteins

Graduate student Rohan Jonnalagadda analyzes the 3D shapes of iron-containing enzymes to parse their role in cellular processes.
Raleigh McElvery
Raised in a computer-savvy family well-versed in software and information technology, Rohan Jonnalagadda had a strong desire to “decode” the world around him. But his kind of code, the genetic one, consists of four repeating letters: A, T, C, and G. “Just like a computer runs on software, I wanted to investigate the code behind the molecular hardware that gives rise to life,” he says. Now a sixth-year graduate student in the Drennan lab, he works to decrypt the structure of metal-containing proteins, in order to determine the roles they play in vital cellular reactions.
When Jonnalagadda was an undergraduate biochemistry major at the University of California, Berkeley, it became clear to him that the genetic code was more than just a string of letters; it also serves as the blueprint for all the proteins in the entire organism. These proteins fold into complex 3D structures, which ultimately beget function.
At UC Berkeley, he joined a lab studying the iron-containing protein Heme-Nitric Oxide/Oxygen (H-NOX) that senses nitric oxide gas in bacterial and eukaryotic cells. When H-NOX binds to nitric oxide, it must change its 3D shape in the process. Jonnalagadda used a technique known as X-ray crystallography to freeze H-NOX in various stages of this conformational change to determine how it binds the gas molecules.
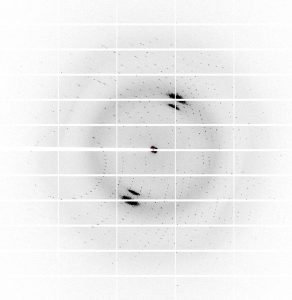

By the time he graduated and began his PhD at MIT Biology, Jonnalagadda had been using X-ray crystallography for over two years. Today, as a member of Catherine Drennan’s lab, he continues to leverage this same method to parse the structure of additional metal-containing proteins.
“I think we sometimes ignore the fact that we need trace metals in order to survive,” he says. “I was interested in continuing to think about what different metals could do in the cell. And using metals opens up a whole new world of chemical reactions that you generally don’t learn about in class.”
In fact, the two projects that he’s devoted most of his time to over the past five years involve reactions that he’d never even heard of before he arrived at MIT. The focus of his first undertaking was the iron-containing enzyme ribonucleotide reductase (RNR), which helps generate deoxyribonucleotides, the building blocks of DNA.
Jonnalagadda aims to understand how this enzyme is regulated to ensure the cell maintains the proper amount of each type of deoxyribonucleotide, in order to properly replicate and repair its genome. If those ratios are incorrect, the cell could experience detrimental stress.
Because the enzyme is regulated differently in humans than it is in bacteria, scientists hope to one day create antibiotics that target the bacterial RNR while leaving the human RNR unscathed. Jonnalagadda works with the human version, devising an assay that will allow him to better assess the differences between the two enzymes. RNR is notoriously difficult to work with, and so Jonnalagadda has spent much of his time developing ways to purify it so it remains stable.
His second project is a collaboration with researchers at his alma mater, UC Berkeley, investigating isonitriles — compounds containing a carbon atom tripled bonded to a nitrogen atom. Because isonitriles are used to make drugs like antibiotics, scientists have a keen interest in exploring new ways to produce them. The team discovered that one bacterium, Streptomyces coeruleorubidus, had a novel and mysterious way of synthesizing these compounds. Jonnalagadda wants to know exactly how these particular bacteria do it.
He is using X-ray crystallography to determine the structure of the iron-containing enzyme ScoE in S. coeruleorubidus, which is responsible for forming the carbon-nitrogen triple bond characteristic of isonitriles.
“It’s exciting to be working on a protein that’s only just been discovered,” he says. “There’s just so much more to learn about its fundamental biological function. I think that’s why basic research is so appealing to me; you never know where the work will take you, or the impacts it could have on human health later on.”
Extending the frontiers of any discipline requires some guesswork and metaphorical bushwhacking, and Jonnalagadda has learned almost as much from his failed experiments as he has from his successful ones. “I’m proud that I’ve been able to use what I’ve learned about experimental design to help others in my lab when they have questions,” he says.
As he considers life post-graduation, he hopes to use the biochemical and structural techniques he’s mastered over the years to secure a job in industry.
“Being part of a department with such broad and wide-ranging research interests has made it easy to see that my work doesn’t exist in a vacuum,” he says. “It connects to many different aspects of biology.”
