Junk DNA makes a comeback

Third-year graduate student Emma Kowal is searching DNA for sequences that regulate gene expression.
Saima Sidik
“I went into science because of a certain obsession with the romance of it,” says Emma Kowal, a third-year graduate student in Chris Burge’s lab in the MIT Department of Biology. “I loved the idea of the scientist as an adventurer exploring the frontiers of knowledge and the universe. And I haven’t let go of that yet.”
Kowal has always been an avid science fiction reader, and now she’s living out a real-life scientific odyssey. The quest she’s taken on for her PhD research involves an understudied type of DNA sequence called an intron, and the roles that introns might play in regulating gene expression.
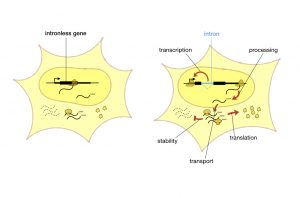
Introns lie between the DNA sequences that cells use for protein production, and are initially incorporated into the messenger RNA, or mRNA, that cells produce as an intermediate step in synthesizing proteins from DNA. But before they complete protein synthesis, cells remove introns from mRNA through a process called splicing, which has led many people to view introns as junk DNA with splicing acting like a garbage disposal.
“Introns appeal to me as the underdog genomic region,” Kowal says. Although they’re often seen as unimportant, introns are ubiquitous and plentiful, collectively making up 24% of the human genome. All eukaryotes have them, and, on average, each human gene encodes eight. Many researchers, including Kowal, think that introns have been underestimated, and that they may play an important role in regulating gene expression.
Introns are only the latest chapter in Kowal’s RNA story. She began her research career as a Harvard University undergraduate student working in the Szostak Lab at Massachusetts General Hospital, where she studied how RNA catalyzed the evolution of cells on the early earth. Although studying primordial life was intellectually stimulating, Kowal wanted to work on something more applied, and so she joined the Church Lab in the Harvard Department of Genetics. There she developed methods for purifying and imaging enigmatic RNA-containing lipid compartments called extracellular vesicles, which cells release into their surrounding environments possibly to communicate with one another.

For the sequel to her bachelor’s degree, Kowal chose to attend MIT Biology because she’d heard that, “at MIT, everyone is one standard deviation nerdier, on average, than they are at other schools.” In this sense, she has not been disappointed. Kowal calls the energy at MIT “unparalleled,” and she says, “people are jazzed about what they’re doing, and the whole campus reflects that.”
In some ways, these reflections are physical. Much of the artwork around MIT pays homage to major scientific discoveries, and Kowal says this reverence for science is one factor that attracted her to MIT. From the mural of DNA in the Biology Department to the golden neurons that descend alongside the staircase in the McGovern Institute for Brain Research, it’s as if the community is saying, “look at how awesome the universe is!” as Kowal puts it.
In other ways, this energy is reflected in the people she converses with daily. “I really like the students here,” Kowal says. “Everyone is enthusiastic, but also down to earth.” When she’s not exploring the realms of science, Kowal sometimes has more fanciful adventures with the Dungeons and Dragons group that she’s formed with some of her classmates.
Kowal didn’t necessarily intend to continue working on RNA at MIT Biology, but when she heard about Chris Burge’s lab, which focuses on RNA and the proteins that mediate its production and stability, she felt a call to action.
The Burge Lab combines high throughput experimental techniques with bioinformatics, and Kowal wants to develop expertise in both these fields. “If you’re skilled in generating and analyzing big data sets, you can ask questions that other people can’t,” she says. The Burge lab seemed like the perfect setting for her PhD.
Burge asks his students to begin their degrees with a month-long reading period during which they sift through the literature to find a topic that they want to study. “You’re not allowed to pick up a pipette or do any analysis during your reading period,” Kowal says. “You just read and discuss your ideas and let things percolate.” As she read, Kowal came across a number of studies that discussed the influence that introns have on gene expression levels.
Over and over, scientists have noticed that cells produce more protein from genes that contain introns than when those same introns are removed. Intron mediated enhancement (IME), as this effect is called, is a “stunningly broad phenomenon,” Kowal says, and scientists have observed it in a wide range of organisms, from yeast to plants to humans.
Splicing machinery, which removes introns from mRNA, likely plays a role in IME. This machinery binds mRNA as it’s being produced from DNA, then interacts with, and influences, the RNA production machinery. However, researchers have created mutant introns that can’t be recognized by splicing machinery, and sometimes these introns still enhance gene expression, so splicing isn’t the only factor that drives IME. Moreover, replacing one intron with another of the same size containing a different DNA sequence can change its effect, implying that the exact DNA sequences within introns may dictate their effects on gene expression. Kowal is intrigued by this last point, and wants to find these intronic sequences and figure out which have the largest effects on gene expression and why.
“This is an old mystery that’s ripe for new tools,” Kowal says. Over the last decade, researchers have begun using a technique called RNAseq to count the copies of mRNA that are made from each gene in a population of cells. Instead of replacing an intron with a single alternative DNA sequence, Kowal plans to replace an intron with a myriad of random DNA sequences, then use RNAseq to count how many copies of mRNA cells make when they encode each of these random introns.
Preparing to test these random sequences has been an odyssey in and of itself, and Kowal has spent the last year building the system that she’ll use. First, she needed to decide which intron to replace. She chose one from a gene called UbC. Removing this intron reduces expression of UbC by ten-fold.
Besides contributing strongly to IME, the UbC intron is a great candidate for Kowal’s experiment because it lies in a regulatory region of the UbC mRNA that precedes the portion that’s translated into protein. This let her replace the UbC protein coding region with a fluorescent protein that she’ll use to visualize how much protein cells make when they encode each random intron sequence.

Kowal has spent the last year meticulously incorporating a library of random introns into this synthetic version of the UbC gene. She anticipates being able to introduce them into cells soon, to see which random introns result in the highest levels of mRNA and protein production. Thanks to RNAseq, she’ll be able to monitor how much each random intron contributes to mRNA expression. Because she can measure how brightly the fluorescent protein glows, she can correlate these mRNA levels with protein levels. From this, she’ll learn which intron sequences enhance gene expression most strongly, and she’ll also know whether these introns lead to higher levels of mRNA production, or if the same amount of mRNA is made into more protein. This distinction will offer her clues about the mechanism that introns use to enhance gene expression.
Once Kowal knows which intron sequences promote gene expression most effectively, she’ll take advantage of the Burge lab’s bioinformatics expertise to analyze the distribution of these sequences throughout genomes and predict how they affect global gene expression. Kowal suspects certain intron sequences are bound by proteins that mediate mRNA production and stability, and she thinks her work will identify these protein-intron pairs.
Kowal balances her scientific adventures with outdoor adventures. Specifically, she’s recently fallen in love with rock climbing. “Climbing is a great counterpart to science because it’s something you can chip away at, and then there’s this huge satisfaction when you finally achieve a climb,” she says. “And also, between climbing and pipetting, I have really strong fingers.”
As for her love of science fiction, Kowal hopes to one day pen a science-based adventure of her own, but not before she’s made her mark as a scientist, either as a professor or in industry. ”It makes sense for me to focus most of my energy on science right now,” she says. “But after I’ve led a spectacular, adventurous life in science, maybe I’ll use my reflections to write a novel.”
