
Raleigh McElvery
May 19, 2021
The DNA inside a single human cell is several meters long — yet it must be condensed to fit inside a space one-tenth the diameter of a hair. That’s like stretching a string from Philadelphia, Pennsylvania to Washington, D.C., and then trying to stuff it into a soccer ball. Imagine then organizing all of this information for each of the body’s 3 trillion cells! The DNA is condensed by proteins called histones that create a spool around which the DNA can wrap itself. How tightly the DNA is wound determines whether it is accessible enough for other proteins to bind to and copy into RNA, toggling gene expression levels up or down.
One specialized type of histone, H2A.Z, is ubiquitous and essential among multicellular organisms. But there have been conflicting reports about how it affects gene expression, especially during embryonic development.
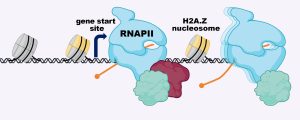
Several years ago, Laurie Boyer’s lab at MIT was the first to show that H2A.Z wraps the DNA located around the start sites of most genes, where the molecular machine RNA polymerase II (RNAPII) binds to copy the DNA into RNA. Boyer’s team demonstrated that removing H2A.Z prevented embryonic cells from turning on genes that are important for forming organs and tissues. But scientists still weren’t sure how H2A.Z exerted its effects.
Now, in a recent Nature Structural and Molecular Biology study, a team from the Boyer lab, led by former postdoc Constantine Mylonas, has revealed how H2A.Z regulates the ability of RNAPII to properly transcribe DNA into the messages that specify all cell types in the body. The researchers found that in embryonic stem cells, H2A.Z serves as a “yellow traffic light,” signaling RNAPII to slow the process of transcribing DNA into RNA. Although there are other proteins that also contribute to RNAPII pausing, H2A.Z establishes a second barrier to transcription that allows gene expression to be tuned in response to developmental signals.
“H2A.Z appears to regulate how fast RNAPII begins to transcribe DNA, and this allows the cell time to respond to important cues that ultimately direct a stem cell to become a brain or heart cell, for example,” says Boyer, a professor of biology and biological engineering. “This connection was a critical missing piece of the puzzle, and explains why H2A.Z is essential for development across all multicellular organisms.”
According to Boyer, H2A.Z’s role in gene expression has been difficult to pin down because previous approaches only provided static snapshots of how proteins interact with DNA days after loss of the histone. Boyer’s team overcame this shortcoming by leveraging a system that allowed for targeted degradation of H2A.Z within hours. They combined this technique with high-resolution genomic approaches and live cell imaging of RNAPII dynamics using super-resolution microscopy. With help from Ibrahim Cissé’s lab, they were able to visualize RNAPII dynamics in real time at the single molecule level in embryonic stem cells. Upon loss of H2A.Z, they found a remarkable increase in RNAPII movement in the cells, consistent with their genomic results showing a faster release of RNAPII and an increase in transcription in the absence of H2A.Z.
Next, the researchers plan to determine precisely how H2A.Z is targeted to the start sites of genes and how it forms a barrier to RNAPII passage.
Boyer says pinpointing the way histone variants like H2A.Z control gene expression is fundamental to understanding how developmental decisions are made, and will help researchers understand why misregulation of H2A.Z has been linked to diseases such as cancer.
“Emerging evidence indicates that DNA ‘packaging proteins’ like histones directly participate in how RNAPII can read and transcribe DNA,” she explains, “and that crucial connection wasn’t clear before.”
Image credits: courtesy of Laurie Boyer
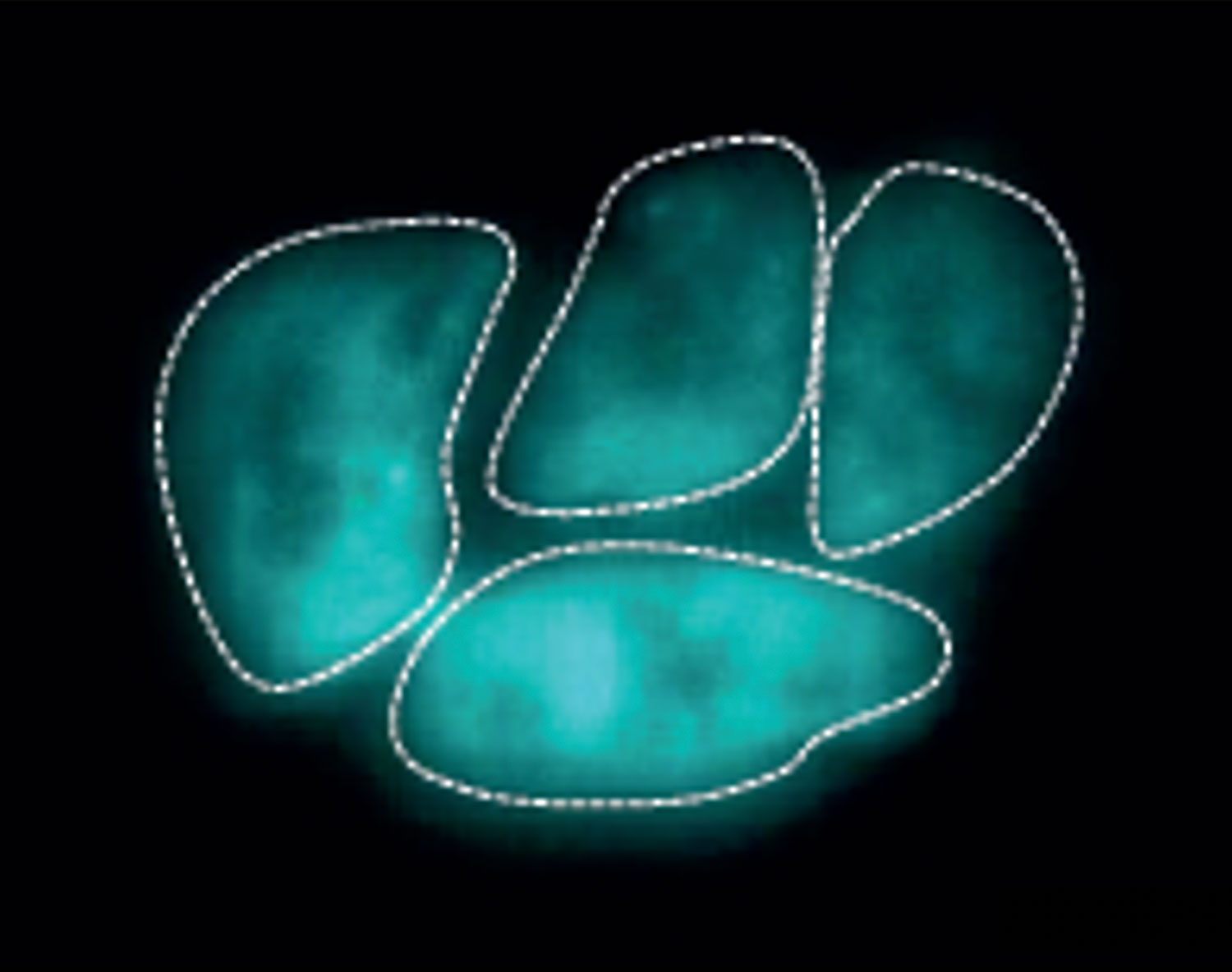
Top image: Live cell super-resolution imaging showing RNAPII dynamics at a single molecule level in embryonic stem cells. The bright and colored clusters represent RNAPII molecules.
Citation:
“A dual role for H2A. Z. 1 in modulating the dynamics of RNA Polymerase II initiation and elongation.”
Nature Structural & Molecular Biology, online May 10, 2021, DOI: 10.1038/s41594-021-00589-3
Constantine Mylonas, Choongman Lee, Alexander L. Auld, Ibrahim I. Cisse, and Laurie A. Boyer.
