New finding suggests differences in how humans and bacteria control production of DNA’s building blocks.
Anne Trafton | MIT News Office
February 20, 2018
Using a state-of-the-art type of electron microscopy, an MIT-led team has discovered the structure of an enzyme that is crucial for maintaining an adequate supply of DNA building blocks in human cells.
Their new structure also reveals the likely mechanism for how cells regulate the enzyme, known as ribonucleotide reductase (RNR). Significantly, the mechanism appears to differ from that of the bacterial version of the enzyme, suggesting that it could be possible to design antibiotics that selectively block the bacterial enzyme.
“People have been trying to figure out whether there is something different enough that you could inhibit bacterial enzymes and not the human version,” says Catherine Drennan, an MIT professor of chemistry and biology and a Howard Hughes Medical Institute Investigator. “By considering these key enzymes and figuring out what are the differences and similarities, we can see if there’s anything in the bacterial enzyme that could be targeted with small-molecule drugs.”
Drennan is one of the senior authors of the study, which appears in the Feb. 20 issue of the journal eLife. JoAnne Stubbe, the Novartis Professor of Chemistry Emerita at MIT, and Francisco Asturias, an associate professor of biochemistry at the University of Colorado School of Medicine, are also senior authors. The paper’s lead authors are MIT research scientist Edward Brignole and former Scripps Research Institute postdoc Kuang-Lei Tsai, who is now an assistant professor at the University of Texas Houston Medical Center.
An unusual enzyme
The RNR enzyme, which is found in all living cells, converts ribonucleotides (the building blocks of RNA) to deoxyribonucleotides (the building blocks of DNA). Cells must keep a sufficient stockpile of these building blocks, but when they accumulate too many, RNR is shut off by a deoxynucleotide molecule known as dATP. When more deoxynucleotides are needed, a related molecule called ATP binds to RNR and turns it back on.
An unusual feature of RNR is that it can catalyze the production of four different products: the nucleotide bases often abbreviated as A, G, C, and T. In 2016, Drennan discovered that the enzyme achieves this by changing its shape in response to regulatory molecules.
Most of the researchers’ previous work on RNR structure has focused on the version found in E. coli. For those studies, they used X-ray crystallography, a technique that can reveal the atomic and molecular structure of a protein after it has been crystallized.
In the new study, Drennan and her colleagues set out to examine the human version of RNR. This protein’s structure, which turned out to be very different from the bacterial version, proved elusive using X-ray crystallography, which doesn’t work well for proteins that don’t readily crystallize. Instead, the researchers turned to an advanced form of microscopy known as cryo-electron microscopy (cryo-EM).
Until recently, cryo-EM typically offered resolution of about 10 to 20 angstroms, which might reveal the overall shape of a protein but no detail about the position and shape of smaller structural units within it. However, in the past few years, technological advances have led to an explosion in the number of structures achieving resolutions of about 3 angstroms. That is high enough to trace individual protein chains within the larger molecule, as well as internal structures such as helices and even side chains of amino acids.
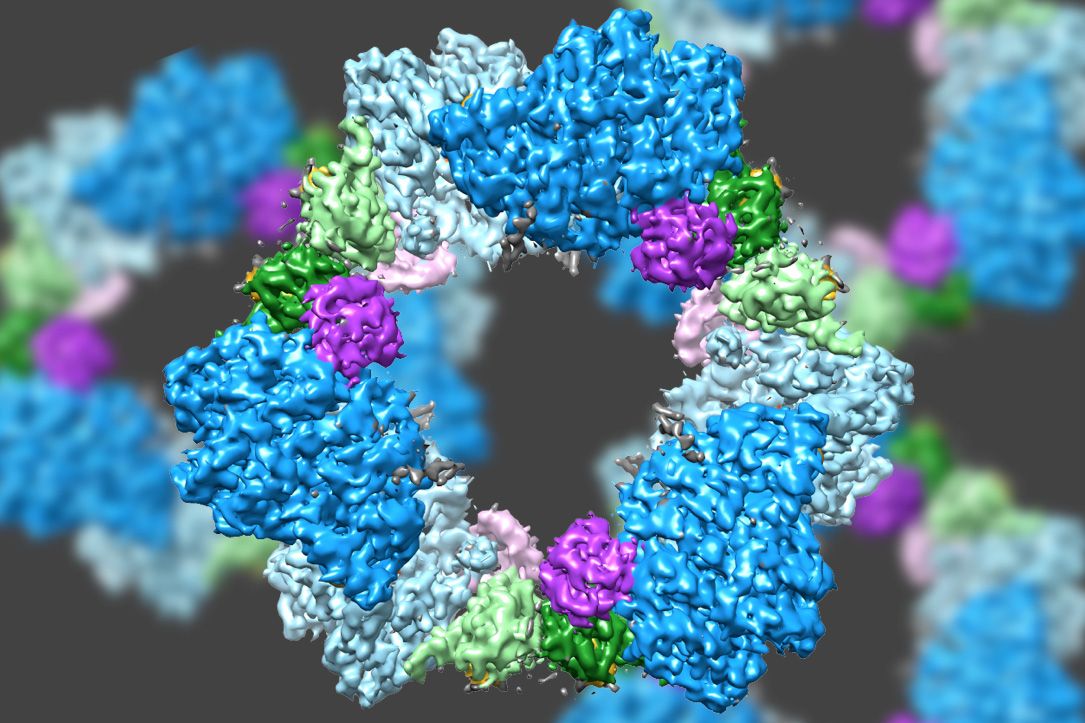
Scientists already knew that RNR consists of two protein subunits known as alpha and beta. Using cryo-EM, the MIT team found that the human version of the enzyme forms a ring made from six of the alpha subunits. When ATP, which activates RNR, is bound to the enzyme, the ring is unstable and can be easily opened up, allowing the beta subunit to make its way into the ring. This joining of alpha and beta allows the enzyme’s active site, located in the beta subunit, to perform the chemical reactions necessary to produce deoxynucleotides.
However, when the inhibitor dATP is present, the ring becomes much more rigid and does not allow the beta subunit to enter. This prevents the enzyme from catalyzing the production of deoxynucleotides.
Designing drugs
Several cancer drugs now in use or in development target the human version of RNR, interfering with cancer cells’ ability to reproduce by limiting their supply of DNA building blocks. The MIT team has found evidence that at least one of these drugs, clofarabine diphosphate, works by inducing the formation of rigid 6-unit alpha rings.
This 6-unit ring is not found in the bacterial form of RNR, which instead assembles into a distinct ring containing four alpha subunits and four beta subunits. This means it could be possible to design antibiotics that target the bacterial version but not the human version, Drennan says.
She now plans to investigate the structures of other protein molecules that are difficult to study with X-ray crystallography, including proteins with iron sulfur clusters, which are found in many metabolic pathways. The microscopy work in this study was performed at the Scripps Research Institute, but when MIT’s new MIT.nano building opens, it will house two cryo-EM microscopes that will be available to the MIT community as well as other potential users in industry and academia.
“The technological advances that have allowed cryo-EM to get to such high resolution are really exciting,” Drennan says. “It’s really starting to revolutionize the study of biology.”
The research was funded by the National Institutes of Health.
