Firefly genomics reveals independent evolution of bioluminescence in beetles
Lisa Girard | Whitehead Institute
October 16, 2018
Cambridge, MA — Researchers at Whitehead Institute and collaborators from fourteen other institutions around the world have shed light on the evolutionary origins of luciferase, the key enzyme behind the glow of fireflies and other bioluminescent beetles. By sequencing the genomes of two American and Japanese firefly species that diverged approximately 100 million years ago, along with a more evolutionarily distant bioluminescent Caribbean click beetle, the team discovered that luciferase appears to have arisen independently in fireflies and click beetles. Examining the genes flanking that encoding the luciferase gene, suggests an evolutionary path along which the luciferase gene arose from duplications and divergences of CoA ligase genes involved in fat metabolism. As described online October 16 in the journal eLife, these findings provide fundamental insights into how enzymes can evolve, potentially inform strategies to help protect bioluminescent beetles from a shifting climate and habitat, and could extend the utility of luciferase, which has also been harnessed for biomedical and agricultural research, as a laboratory tool.
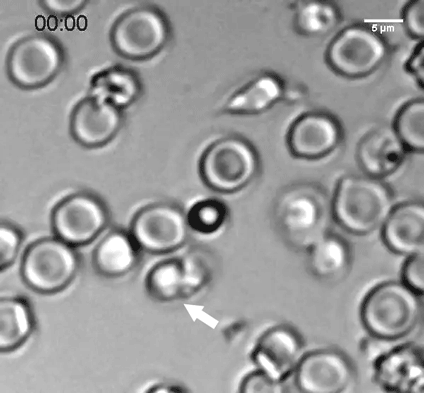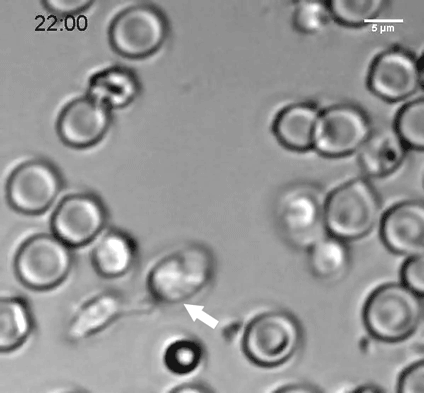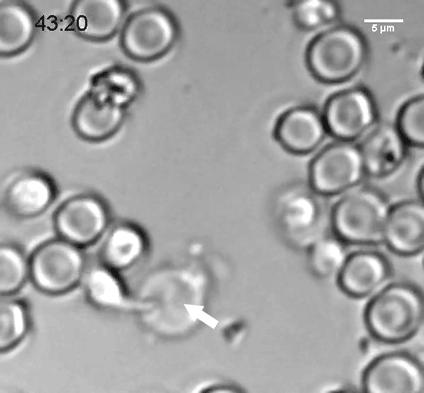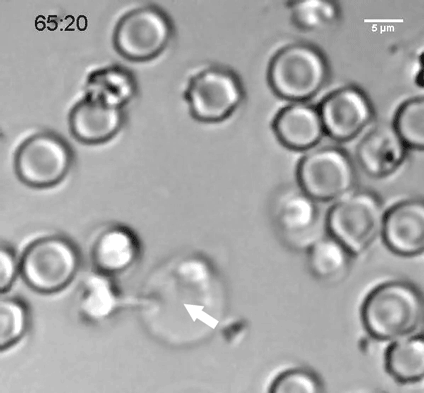
Throughout much of the world, the silent flash of a firefly on a warm evening can only mean one thing-Summer has arrived. But fireflies don’t just signal summer, their glow serves as a mating signal to other fireflies, and is even a warning that they are chemically defended, having a noxious taste capable of repelling the boldest of predators.
Belying its grandeur, the chemistry of firefly bioluminescence is relatively straightforward. Their light is produced by a specialized firefly enzyme, luciferase, that breaks down a molecule called luciferin, producing light in the process. Luciferase has become a mainstay tool in the laboratory. Scientists can fuse their gene of interest to luciferase and assay for gene expression by measuring the intensity of the glow after luciferin is added.
Beyond fireflies, there are other bioluminescent beetles (despite their name, fireflies are actually beetles), including certain tropical click-beetles. Perplexingly, these diverse bioluminescent beetles use very similar luciferase enzymes and luciferin molecules, but have an unrelated anatomy of their light-producing organs (also known as lanterns), making it unclear if their bioluminescence evolved from a common luminous ancestor, or if their special glow evolved independently.
Since fireflies and bioluminescent click-beetles are not model organisms like mice or fruit flies for which there is a wealth of genetic information, Jing-Ke Weng, Whitehead Institute Member and assistant professor of biology at Massachusetts Institute of Technology (MIT), along with a graduate student in Weng’s lab, Tim Fallon, and Cornell postdoctoral researcher, Sarah Lower, began their investigations by sequencing the genome of the American Big Dipper firefly, Photinus pyralis. Named for its distinctive swooping “J” flash , this common inhabitant of meadows and suburban lawns has been called the “All-American firefly”. Due to its abundance and ease of identification, it was also the firefly of choice for scientific study, and is the species from which luciferin and luciferase were first characterized. Wanting to start their work quickly and make their progress and data available to others in the firefly community, Whitehead Institute researchers and collaborators crowdsourced funds to sequence the Big Dipper firefly.
The Big Dipper genome sequence, they discovered, revealed interesting insights into the origin of the luciferase gene. Examining the genes flanking that encoding luciferase, they found a cluster, or tandem repeat, of fatty acid CoA ligase genes with the luciferase gene sitting in the middle of this cluster. Sequence similarity and proximity between the luciferase and fatty acid CoA ligase genes suggested an evolutionary path along which the luciferase gene was produced from tandem duplication and divergence of an ancestral fatty acid CoA ligase gene.
“When the luciferase gene was cloned, people knew it was similar to the fatty acid CoA ligase gene in sequence, and hypothesized that it must be related to that ancestry. But what we uncovered from the luciferase gene locus is a tandem repeat of five genes, four are still the fatty acid CoA ligases, but then luciferase evolved right in the middle we believe from divergence of one of these duplications,” says Weng.
The Big Dipper sequencing provided important insights into the origin of luciferase and additional factors involved in bioluminescence, but in order to gain additional insights into the evolution of bioluminescence, the researchers set out to sequence two additional species that they hoped would provide the additional context to help them triangulate on some answers.
The bioluminescent click beetle, Ignelater luminosus, is related to the firefly, but on another branch of the tree of life entirely. Instead of producing light at its tail, it has two lanterns behind its head.
“We thought that sequencing the click beetle would provide insights into the evolution of bioluminescence as well as perhaps into how these animals could acquire very similar traits in terms of their biochemistry, but not in terms of their development,” says Fallon.
The third species they selected to sequence was a Japanese aquatic firefly (Aquatica lateralis), known in Japan as the Heike firefly. Heike and the Big Dipper diverged from one another over 100 million years ago (to give you a sense of how far this is, it is older than the evolutionary distance between humans and rodents).
The researchers analyzed genomic data from the Japanese aquatic firefly and saw a similar arrangement around the luciferase gene locus as they had in the Big Dipper genome, suggesting that luciferase arose from a common ancestral event in both firefly species. The structure around the luciferase locus in the click beetle, however, was entirely absent, suggesting that luciferase arose through a different event. Taken together, by sequencing and analyzing data from the genomes of two firefly species that diverged approximately 100 million years ago, along with a more evolutionarily distant bioluminescent click beetle, the team discovered that luciferase appears to have evolved independently in both fireflies and click beetles.
“Having the genome allowed us to understand how the evolution of luciferase happened. Before sequencing, we knew there were five genes cloned, including luciferase, in firefly. By sequencing the genomes we actually uncovered those genomic loci where those initial gene duplication events occured,” says Lower.
In addition to the origins of luciferase, these findings also provided the researchers with insights into the evolution of the light organs.
“Since our findings suggest that luciferase originated independently in both lineages, we can infer that anything that came after luciferase, for example the light organs, or other things dependent on luciferase should also be independent,” says Fallon.
Discovering how bioluminescence arose, as well as other complex traits, can be studied now that genomic information is available. The information can also inform strategies to protect fireflies, whose populations in many parts of the world are diminishing. In addition to adding tools to help reveal a constituent parts list that could allow researchers to optimize bioluminescence as a tool, these findings reveal important insights into the evolution of bioluminescence as well as genomic evolution more broadly.
“Luciferase is a perfect example of how to build a new enzyme, duplication of a related progenitor gene followed by mutation and selection,” says Weng. “And one of the most exciting parts of this study was that by examining the evolutionary scars in the genomes we studied we could actually see it happen.”