As Duke University’s provost since 2014, she has advocated for faculty excellence and reinforced the institution’s commitment to the student experience.
Steve Bradt | MIT News Office
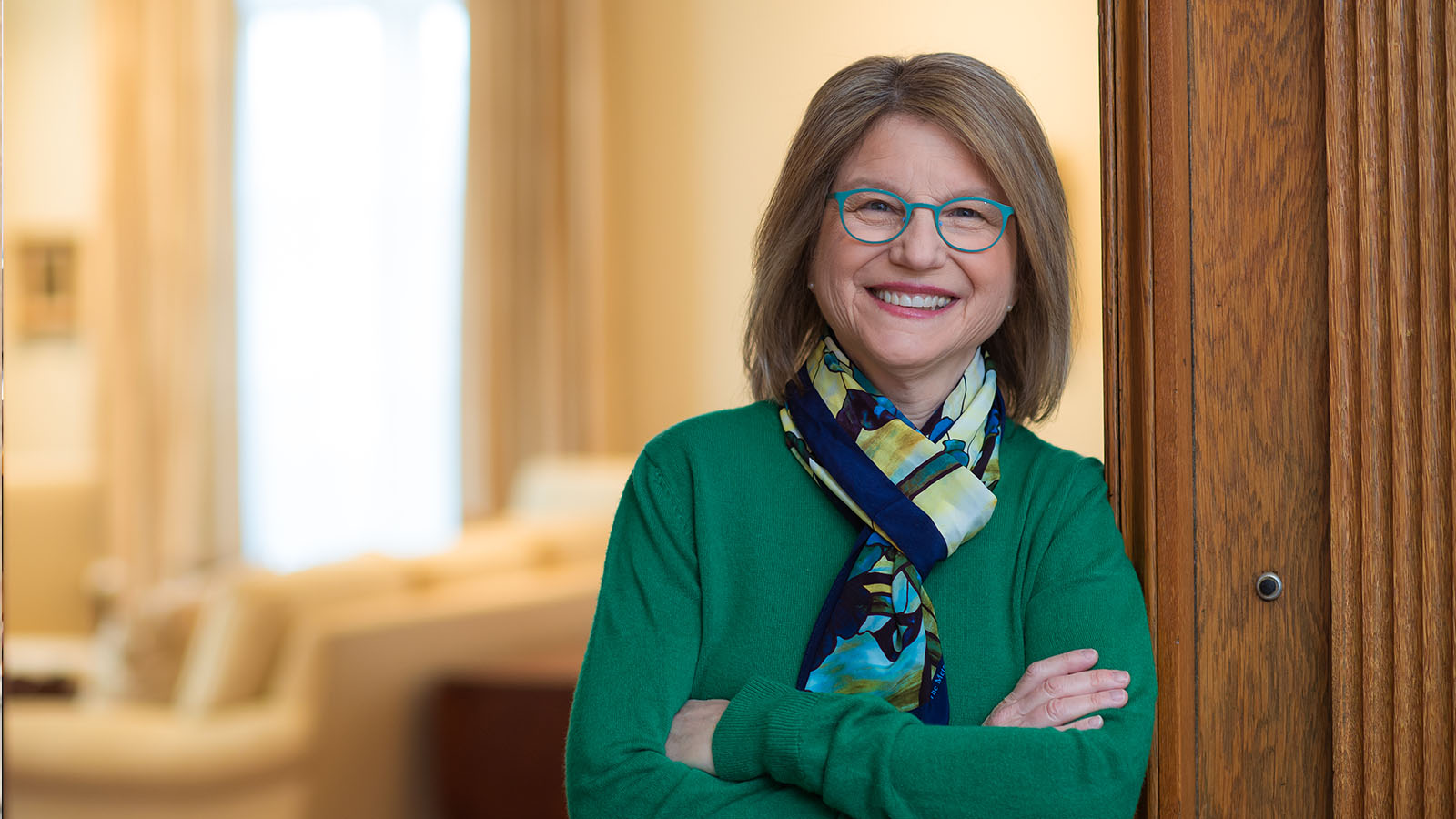
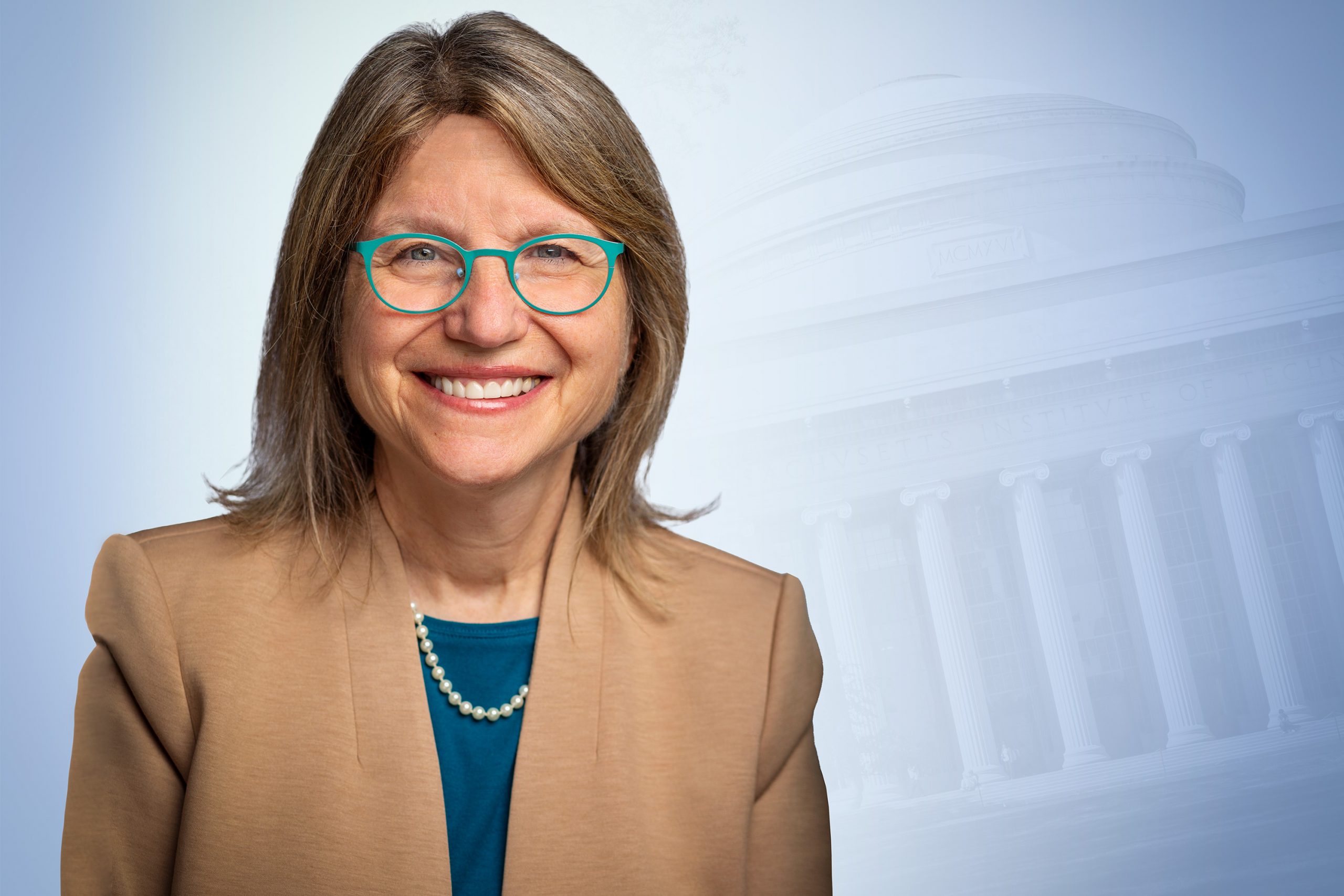
Sally A. Kornbluth, a cell biologist whose eight-year tenure as Duke University’s provost has earned her a reputation as a brilliant administrator, a creative problem-solver, and a leading advocate of academic excellence, has been selected as MIT’s 18th president.
Kornbluth, 61, was elected to the post this morning by a vote of the MIT Corporation. She will assume the MIT presidency on Jan. 1, 2023, succeeding L. Rafael Reif, who last February announced his intention to step down after 10 years leading the Institute.
A distinguished researcher and dedicated mentor, Kornbluth is currently the Jo Rae Wright University Professor of Biology at Duke. She has served on the Duke faculty since 1994, first as a member of the Department of Pharmacology and Cancer Biology in the Duke University School of Medicine and then as a member of the Department of Biology in the Trinity College of Arts and Sciences.
As Duke’s provost since 2014, Kornbluth has served as the chief academic officer of one of the nation’s leading research universities, with broad responsibility for carrying out Duke’s teaching and research missions; developing its intellectual priorities; and partnering with others to achieve wide-ranging gains for the university’s faculty and students. She oversees Duke’s 10 schools and six institutes, and holds ultimate responsibility for admissions, financial aid, libraries, and all other facets of academic and student life.
“The ethos of MIT, where groundbreaking research and education are woven into the DNA of the institution, is thrilling to me,” Kornbluth says. “The primary role of academic leadership is in attracting outstanding scholars and students, and in supporting their important work. And when it comes to the impact of that work, MIT is unparalleled — in the power of its innovations, in its ability to move those innovations into the world, and in its commitment to discovery, creativity, and excellence.”
“My greatest joy as a leader has always been in facilitating and amplifying the work of others,” Kornbluth adds. “I am eager to meet all the brilliant, entrepreneurial people of MIT, and to champion their research, teaching, and learning.”
A broad search with extensive consultation
Kornbluth’s election as MIT’s president is the culmination of an eight-month process in which a 20-member presidential search committee generated a list of approximately 250 possible candidates for the presidency. These candidates brought a broad range of backgrounds in academia and beyond, and included both members of the MIT community and people outside the Institute.
“Dr. Sally Kornbluth is an extraordinary find for MIT,” says MIT Corporation Chair Diane B. Greene SM ’78. “She is decisive and plain-spoken, a powerhouse administrator who has proactively embraced critical issues like free speech and DEI. An accomplished scientist with a liberal arts background, Dr. Kornbluth is a broadly curious, principled leader who deeply understands MIT’s strengths. Her vision and her humanity will inspire our hearts and minds, and her comprehension of the importance of discovery, innovation, and entrepreneurship will serve us well as MIT confronts the challenges of today’s world.”
The presidential search committee was chaired by MIT Corporation Life Member John W. Jarve ’78, SM ’79. Under his leadership, the committee conducted comprehensive outreach with MIT faculty, students, staff, alumni, and individuals beyond MIT.
“Through these exhaustive efforts, we created a list of attributes for MIT’s next president, to ensure our new leader would have a successful tenure at MIT, would be widely embraced by the MIT community, and would maintain MIT’s excellence as the world’s leading science and technology university,” Jarve says. “I am confident that we have found that leader in Sally Kornbluth, who appreciates MIT’s uniqueness, is committed to maintaining its standards of excellence, and is intellectually broad and insatiably curious.”
“Although she is new to MIT, Sally Kornbluth is a scholar who seems cut from our own cloth,” adds Lily L. Tsai, the Ford Professor of Political Science and chair of the MIT faculty, who also served on the search committee. “She is a bold leader with exceptional judgment; an active listener who seeks all viewpoints with a genuinely open-minded approach; a principled, high-integrity individual who is trusted by her community; and a person with experience handling crises with wisdom and calm. I look forward to welcoming her to our community.”
Wide-ranging gains for Duke faculty and students
After becoming Duke’s provost on July 1, 2014, Kornbluth quickly established herself as a transformative leader who partnered eagerly with faculty and others to build upon the university’s strengths. The first woman to serve Duke as its provost, she became a forceful advocate for faculty excellence, advancement, and diversity.
“The presidency of MIT is a wonderful responsibility,” says outgoing President L. Rafael Reif. “Known for her brilliance, wide-ranging curiosity, and collaborative, down-to-earth style, Sally Kornbluth is a terrific choice to lead our distinctive community, and I look forward to seeing MIT continue to flourish under her leadership.”
As provost, Kornbluth prioritized investments to fortify Duke’s faculty, strengthened its leadership in interdisciplinary scholarship and education, and pursued innovations in undergraduate education. She guided the development of a strategic plan, called Together Duke, that engaged faculty from across the university to advance its educational and research mission.
She also spearheaded a concerted effort to cultivate greater strength in science and engineering at Duke, complementing its longstanding prominence in the humanities and social sciences. That effort has led to the addition in recent years of more than two dozen Duke faculty members in the sciences and engineering, with particular focus on quantum computing, data science, materials science, and biological resilience.
Simultaneously, Kornbluth led efforts to develop a pipeline of faculty from underrepresented groups, aiming to make Duke more diverse and inclusive. She created an Office for Faculty Advancement that helped to grow the number of Black faculty members across campus from 67 in 2017 to more than 100 today, and provided seed money for projects aimed at creating a more inclusive environment for underrepresented faculty as well as funding scholarly projects on race and social equity.
As provost, Kornbluth also reinvigorated Duke’s commitment to the student experience, both in and out of the classroom. Her team sought opportunities to make Duke more accessible and affordable, including new scholarships for first-generation students; increases in need-based financial aid; a preorientation program that includes all first-year students; and a new residential system that more closely links living and learning. During her tenure, Duke has also launched university-wide courses that Kornbluth describes as “essential things for every student to understand,” on topics such as race and climate change.
Kornbluth has adapted some of the lessons from those undergraduate-focused initiatives to benefit graduate and professional students, while partnering with Duke’s Graduate School and her vice provosts to improve the quality of mentoring and other support for graduate students.
She oversaw the launch of the undergraduate degree program at Duke Kunshan University, a liberal arts and research university created in partnership with Wuhan University to offer academic programs for students from China and throughout the world. She has sought to extend Duke’s international outreach and has encouraged the development of new partnerships with a focus on social, economic, and environmental issues impacting societies around the world.
Kornbluth also guided many of Duke’s schools, centers, and institutes through significant leadership transitions. She oversaw a number of key leadership hires, including the appointment of new deans for Duke’s Trinity College of Arts and Sciences, the Pratt School of Engineering, Duke Divinity School, the Sanford School of Public Policy, the Nicholas School of the Environment, Duke’s Graduate School, and the Duke University School of Law, as well as the university librarian and a new vice provost for learning innovation and digital education.
“Sally Kornbluth has demonstrated the ability to lead across disciplines, and to catalyze the type of cross-disciplinary initiatives that have been so instrumental to MIT’s ability to contribute advances in technology and engineering for the betterment of the world,” says Kristala L. Jones Prather, the Arthur Dehon Little Professor of Chemical Engineering, who served on the presidential search committee.
From music to political science to genetics
Born in Paterson, New Jersey, Sally Ann Kornbluth grew up in nearby Fair Lawn. Her father, George, was a music-loving accountant; her mother, Myra, was an opera singer who performed regularly at the New York City Opera, the Metropolitan Opera, and elsewhere around the world under the name Marisa Galvany.
Inspired by a high school teacher, Kornbluth studied political science as an undergraduate at Williams College. Early in her undergraduate years, she gave little thought to studying science, until she had to take a course on human biology and social issues as part of distribution requirements needed to graduate.
“I thought it was really interesting, and, once I saw what science was really about, I found it very exciting,” she recalled in a 2014 interview. “I just hadn’t had that opportunity in high school.”
After earning her BA in political science from Williams in 1982, Kornbluth received a scholarship to attend Cambridge University for two years as a Herchel Smith Scholar at Emmanuel College, ultimately earning a BA in genetics from Cambridge in 1984.
Kornbluth returned to the U.S. to pursue a PhD in molecular oncology at Rockefeller University, awarded in 1989, and then went on to postdoctoral training at the University of California at San Diego. She joined the Duke faculty as an assistant professor in the Department of Pharmacology and Cancer Biology in 1994, becoming an associate professor in 2000 and a full professor in 2005.
Research impacts in cellular behavior — and far beyond
At Duke, Kornbluth’s research focused on the biological signals that tell a cell to start dividing or to self-destruct — processes that are key to understanding cancer as well as various degenerative disorders. She has published extensively on cell proliferation and programmed cell death, studying both phenomena in a variety of organisms. Her research has helped to show how cancer cells evade this programmed death, or apoptosis, and how metabolism regulates the cell death process; her work has also clarified the role of apoptosis in regulating the duration of female fertility in vertebrates.
Kornbluth eventually transitioned into administrative roles at Duke for what she describes as “nonaltruistic reasons: I wanted to attract the best possible students, and I wanted better scientific core facilities.” Her first senior administrative position came when she was named vice dean for basic science at the Duke School of Medicine in 2006, a post she held until being named provost in 2014.
In this role, Kornbluth served as a liaison between the dean of medicine and faculty leaders; oversaw biomedical graduate programs; implemented efforts to support research in basic science; allocated laboratory space; oversaw new and existing core laboratories; and worked with department chairs to recruit and retain faculty. From 2009 to 2011, she also oversaw the clinical research enterprise in the Duke School of Medicine.
As Duke’s provost, with a much wider purview, Kornbluth has worked to foster interdisciplinary efforts across campus. “University leaders need to have broad-ranging intellectual curiosity, and interests in a wide range of topics,” she says. “At MIT, I think there is particularly rich potential in the places where science and engineering brush up against the humanities and social sciences. I am eager to soak in the MIT culture, listen, draw out the best from everyone, and do my part to encourage the Institute to grow ever better.”
Members of MIT’s presidential search committee also look forward to Kornbluth’s arrival on campus.
“In our community conversations, we would again and again come back to three important attributes: that the president be someone who embraces MIT’s unique culture, takes care of the people who create it, and is unafraid to improve it,” says committee member Yu Jing Chen ’22, now a graduate student in urban studies and planning. “For these reasons, we couldn’t be more excited to see Sally Kornbluth lead MIT.”
“Sally Kornbluth is someone who cares about people, and she demonstrated at Duke her passion for excellence and her respect for everyone, no matter their role,” says committee member Deborah Liverman, who serves as executive director of MIT Career Advising and Professional Development. “Those are values that are important to the thousands of people who work to keep MIT the extraordinary place it is. I am eager to see what we can accomplish with her leading the way.”
Among other honors, Kornbluth received the Basic Science Research Mentoring Award from the Duke School of Medicine in 2012 and the Distinguished Faculty Award from the Duke Medical Alumni Association in 2013. She is a member of the National Academy of Medicine, the National Academy of Inventors, and the American Academy of Arts and Sciences.
Kornbluth’s husband, Daniel Lew, is the James B. Duke Professor of Pharmacology and Cancer Biology at the Duke School of Medicine. Their son, Alex, is a PhD student in electrical engineering and computer science at MIT, and their daughter, Joey, is a medical student at the University of California at San Francisco.