Study explains why mutations that would seemingly affect all cells lead to face-specific birth defects.
Anne Trafton | MIT News Office
January 24, 2018
About 1 in 750 babies born in the United States has some kind of craniofacial malformation, accounting for about one-third of all birth defects.
Many of these craniofacial disorders arise from mutations of “housekeeping” genes, so called because they are required for basic functions such as building proteins or copying DNA. All cells in the body require these housekeeping genes, so scientists have long wondered why these mutations would produce defects specifically in facial tissues.
Researchers at MIT and Stanford University have now discovered how one such mutation leads to the facial malformations seen in Treacher-Collins Syndrome, a disorder that affects between 1 in 25,000 and 1 in 50,000 babies and produces underdeveloped facial bones, especially in the jaw and cheek.
The team found that embryonic cells that form the face are more sensitive to the mutation because they more readily activate a pathway that induces cell death in response to stress. This pathway is mediated by a protein called p53. The new findings mark the first time that scientists have determined how mutations in housekeeping genes can have tissue-specific effects during embryonic development.
“We were able to narrow down, at the molecular level, how issues with general regulators that are used to make ribosomes in all cells lead to defects in specific cell types,” says Eliezer Calo, an MIT assistant professor of biology and the lead author of the study.
Joanna Wysocka, a professor of chemical and systems biology at Stanford University, is the senior author of the study, which appears in the Jan. 24 online edition of Nature.
From mutation to disease
Treacher-Collins Syndrome is caused by mutations in genes that code for proteins required for the assembly and function of polymerases. These proteins, known as TCOF1, POLR1C, and POLR1D, are responsible for transcribing genes that make up cell organelles called ribosomes. Ribosomes are critical to all cells.
“The question we were trying to understand is, how is it that when all cells in the body need ribosomes to function, mutations in components that are required for making the ribosomes lead to craniofacial disorders? In these conditions, you would expect that all the cell types of the body would be equally affected, but that’s not the case,” Calo says.
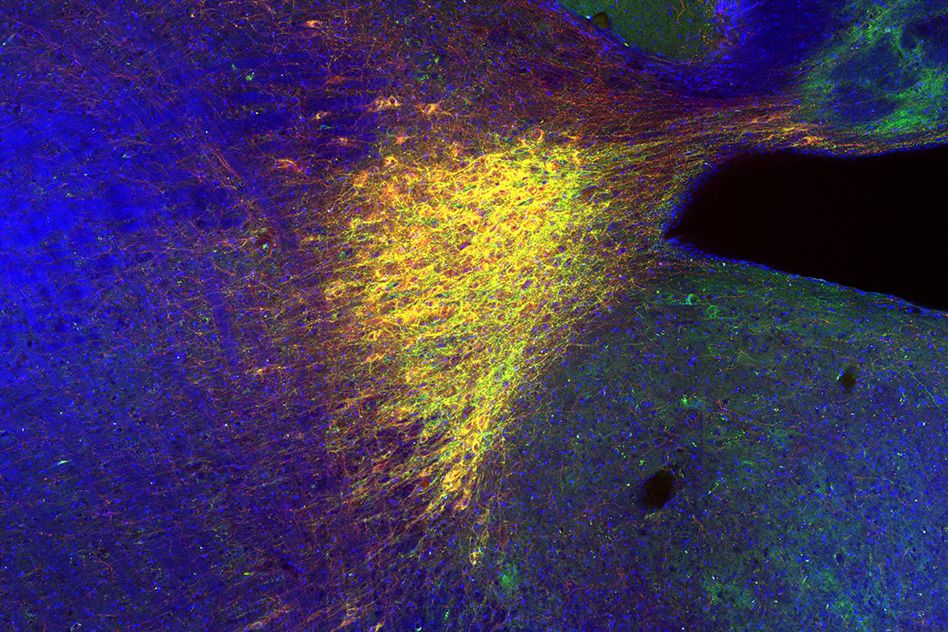
During embryonic development, these mutations specifically affect a type of embryonic cells known as cranial neural crest cells, which form the face. The researchers already knew that the mutations disrupt the formation of ribosomes, but they didn’t know exactly how this happens. To investigate that process, the researchers engineered larvae of zebrafish and of an aquatic frog known as Xenopus to express proteins harboring those mutations.
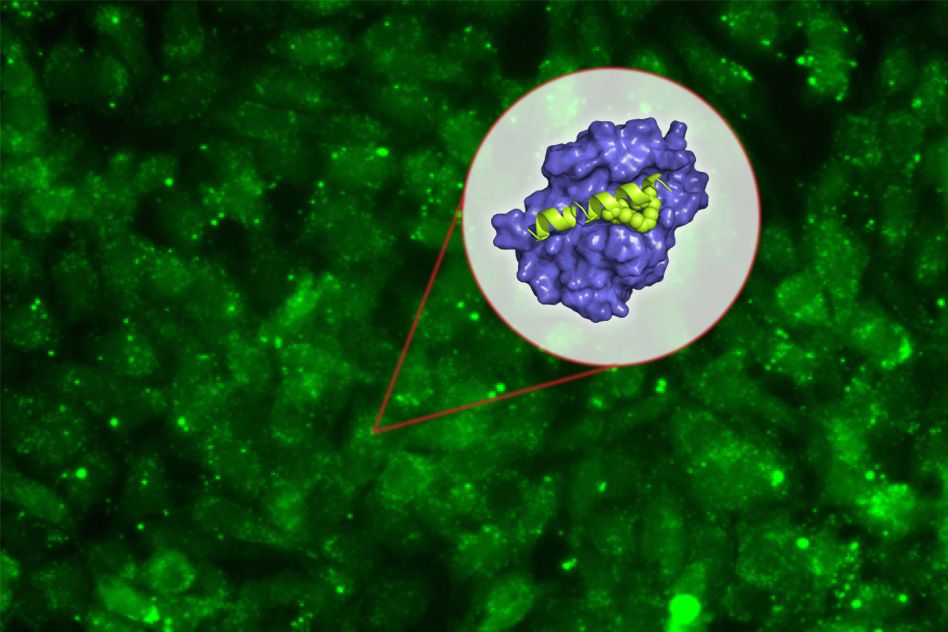
Their experiments revealed that the mutations lead to impairment in the function of an enzyme called DDX21. When DDX21 dissociates from DNA, the genes that encode ribosomal proteins do not get transcribed, so ribosomes are missing key components and can’t function normally. However, this DDX21 loss only appears to happen in cells that are highly sensitive to p53 activation, including cranial neural crest cells. These cells then undergo programmed cell death, which leads to the facial malformations seen in Treacher-Collins Syndrome, Calo says.
Other embryonic cells, including other types of neural crest cells, which form nerves and other parts of the body such as connective tissue, are not affected by the loss of DDX21.
Role of DNA damage
The researchers also found that mutations of POLR1C and POLR1D also cause damage to stretches of DNA that encode some of the RNA molecules that make up ribosomes. The amount of DNA damage correlated closely with the severity of malformations seen in individual larvae, and mutations in POLR1C led to far more DNA damage than mutations in POLR1D. The researchers believe these differences in DNA damage may explain why the severity of Treacher-Collins Syndrome can vary widely among individuals.
Calo’s lab is now studying why affected cells experience greater levels of DNA damage in those particular sequences. The researchers are also looking for compounds that could potentially prevent craniofacial defects by making the cranial neural crest cells more resistant to p53-induced cell death. Such interventions could have a big impact but would have to be targeted very early in embryonic development, as the cranial neural crest cells begin forming the tissue layers that will become the face at about three weeks of development in human embryos.
The research was funded by the National Institutes of Health, Howard Hughes Medical Institute, and March of Dimes Foundation.