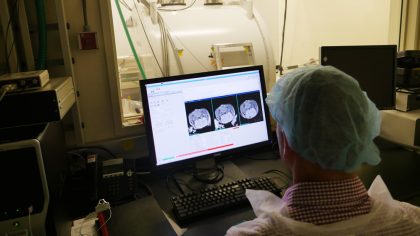
Animal Imaging and Pre-Clinical Testing Core Facility
Koch Institute: Swanson Biotechnology Center
Building 76-188
Contact: Scott Malstrom, PhD
This facility provides access to instrumentation for in vivo, whole animal imaging, including bioluminescence, microCT, PET/CT, ultrasound and small animal MRI technologies. The work here supports translational efforts through the use of refined mouse models of human cancers in preclinical testing of biosensors for tumor detection and monitoring, diagnostic assays, cancer vaccines, drug delivery systems and tumor-targeting modalities, and chemotherapeutic response and drug resistance.

Barbara K. Ostrom (1978) Bioinformatics & Computing Facility
Koch Institute: Swanson Biotechnology Center
Building 76-189B
Contact: Charlie Whittaker, PhD
This facility is a shared institutional resource providing investigators with support, assistance and training in a wide range of bioinformatics topics. The core also maintains a high-performance computing resource offering many bioinformatics applications, substantial processing power and secure storage and provides support for desktop computing and poster printing.

BioMicro Center
Building 68-316
Contact: Stuart Levine
The BioMicro Center provides integrated facilities for high-throughput data-intensive genomics, bioinformatic analysis, as well as large-scale database storage, data mining and data modeling required to fully implement systems approaches to investigate a broad spectrum of biological challenges. Equipment includes Illumina sequencers, Tecan and TTP liquid handlers, microarray scanners, nucleic acid fragment analysis, real-time PCR, plate readers and Luminex technologies, as well as computational servers.

This facility houses instruments to elucidate macromolecular structure, including an analytical ultracentrifuge, a CD spectrometer, a dynamic light scatterer, a differential scanning calorimeter, an isothermal titration calorimeter, a capillary electrophoresis instrument and a bio-layer interferometer. The facility is largely used by researchers in chemistry, biology, and biological engineering, but is available to anyone on campus and beyond.

Biopolymers & Proteomics Core Facility
Building 76-181 (Biopolymers) and 76-185 (Proteomics)
Contact: Dick Cook (Biopolymers) and Amanda Del Rosario (Proteomics)
This facility provides synthetic and analytical capabilities for biological materials, such as proteins, small molecules, and nanoparticles. It encompasses a range of technical expertise and instrumentation including peptide synthesis and peptide arrays; mass spectrometry-based proteomics approaches for identification, characterization, or quantitation of proteins from simple to complex mixtures; mass spectrometry-based mass measurement, mass mapping, and metabolite quantitation; and HPLC analysis and purification.

Center for Automated Cryogenic Electron Microscopy
Building 12, Basement Level
Contact: Sarah Sterling
The Cryo-EM Center is located in MIT.nano, a new shared research building on campus. The facility is commissioning Titan Krios and Talos Arctica cryo-TEMs for high-throughput imaging of purified macromolecules and tomography of cellular specimens. It also includes an Aquilos cryo-FIB/SEM and specimen preparation instrumentation including a Vitrobot and a Chameleon (aka Spot-It-On).

ES Cell and Transgenic Facility
Building 76-195
Contact: Noranne Enzer
This facility supports use of mouse models to study human diseases such as cancer, assisting with design, generation, and analysis of mice carrying engineered mutations. Services include traditional or CRISPR/Cas9 gene targeting in ES cells or embryos; coordination of pre-implantation mouse embryo injections; specialized cell and organoid culture including derivation of germline-competent ES lines and ES cell karyotyping; and assistance, training, and reagents for ES cell, organoid, embryo and mouse analysis.

Functional Genomics Platform
Whitehead Institute, Room 337
Contact: Heather Keys
The Functional Genomics Platform works with users within the Whitehead and affiliated communities to design and implement large-scale pooled CRISPR-Cas9 screens in mammalian systems.

Genetically Engineered Models (GEM) Core Facility
Whitehead Institute, Room 461
Contact: Styliani (Stella) Markoulaki
This facility generates genetically modified mice, using traditional and CRISPR/Cas9 techniques to understand gene function and model human disease. Even though the traditional way is complex, costly and time-consuming, it is still used to achieve difficult mutations. CRISPR in mouse embryos can generate mutations in multiple genes and achieve deletions or insertions in as little as four weeks. Regardless of the method, the facility is here to generate any mouse model in the fastest and most cost-efficient manner.

Genome Technology Core (GTC)
Whitehead Institute, Room 325
Contact: GTC team
This facility provides service and support for high data content genomic technologies including Next-Generation Sequencing, Microarrays, and Quantitative PCR. The GTC team offers sample-to-data solutions for gene expression, protein-DNA binding (ChIP) analysis, microRNA expression, and more.

This facility provides consultation and access to automation instrumentation, including an Arrayscan confocal high content screening microscope, Tecan liquid handlers, plate washers, microplate centrifuges, plate reader, and CO2 incubators. Instruments can be used in walk-up or automated modes using robot arms for general screening or plate preparation. Other services include assay development, access to curated small molecule and lentiviral shRNA (RNAi) libraries, and screening services in both BL2 and BL2+ space.

This facility was established to assist investigators in producing quality histological slides. It houses equipment and expertise to produce sections from frozen, paraffin-embedded, and resin-embedded tissues. Services include production of routine hematoxylin and eosin stained slides, immunohistochemical and special stains, and slide scanning.

iPS Core Facility
Building 46-1303
Contact: Tak Ko
This facility integrates the various research goals of members of the Picower and McGovern Institutes, as well as the Department of Brain and Cognitive Sciences, to create human and animal cell models of diseases. The iPS Core Facility has produced over 60 patient-specific iPS cells from Schizophrenia, Bipolar disease, Depression, Rett syndrome, Alzheimer disease, Down Syndrome, and healthy person’s skin fibroblasts as controls.
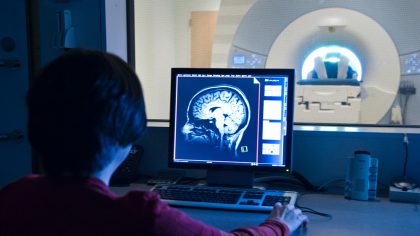
Martinos Imaging Center
Building 46-1171
Contact: MRI Team
This facility, housed within MIT’s McGovern Institute, offers state-of-the-art brain imaging technologies—including human magnetic resonance imaging (MRI), small-animal MRI, electroencephalography (EEG) and magnetoencephalography (MEG)—to MIT researchers and their collaborators throughout the Boston area. Major research themes include: brain mechanisms of perception, memory and emotion; developmental disorders in children; and the neural basis of psychiatric and neurological disorders.

Metabolite Profiling Core Facility
Whitehead Institute, Room 647
Contact: Metabolomics Team
This facility was established to support research related to the role of small molecule and lipid metabolites in cancer biology, aging, and microbial pathogenesis. Facility staff members assist researchers with experimental design and sample preparation strategies, perform data analysis, in addition to developing new analytical techniques for metabolites that are not adequately measured by our existing methods.

Microscopy Core Facility
Building 76-281
Contact: Jeffery Kuhn
This facility provides consultation, assisted imaging, training, and access to imaging platforms, image acquisition equipment and data analysis software. Imaging capabilities include light, epifluorescence, deconvolution and 3D structured illumination microscopy; spinning disk, scanning and multiphoton scanning confocal microscopy; intravital imaging; TIRF, FRAP, FRET and photo-recovery microscopy; fluorescent in situ hybridization; and laser capture microdissection.

Nanowell Cytometry
Building 76-273
Contact: Shanu Mehta
This subgroup in the Flow Cytometry Core Facility supports nanowell microengraving and imaging platforms, enabling the measurement of multiple traits from individual cells derived from a wide array of tissues or cell types, such as immune cells either in their native condition or upon stimulation. The modularity of the platform allows simultaneous assessment from small or large numbers of cells. Additionally, it is possible to obtain genetic information from the individual cells of interest.

Peterson (1957) Nanotechnology Materials Core Facility
Building 76-037
Contact: Abigail Lytton-Jean, PhD
The facility houses a state-of-the-art cryo imaging suite for bio relevant materials, including cryo-transmission electron microscope (TEM), cryo-scanning electron microscope (SEM/FIB) with focused ion beam milling for 3D reconstruction, freeze fracture, freeze substitution, high pressure freezer, and an atomic force microscope (AFM). Instrumentation for material characterization includes high throughput particle sizing (DLS) and counting (Spectradyne), HPLC, dynamic mechanical analysis (DMA), and a rheometer.

This facility provides protein characterization resources utilizing mass spectrometry and liquid chromatography for protein identification, comparative proteomics (using label free, or isotopic labeling (iTRAQ/SILAC)), quantitative proteomics, and the identification of post-translational modifications.

Structural Biology Core Facility
Building 68-588
Contact: Daniel Cham-Chin Lim
This facility provides access to state-of-the-art equipment for X-ray crystallography — including ARI Phoenix and Formulatrix NT8 drop-setting robots for preparation of high-throughput screens for crystallization conditions and RockImager systems at 18°C and 4°C for imaging of the screening experiments. There is also a rotating anode X-ray generator for in-house data collection, and advice can be provided regarding optimizing crystallization conditions, processing diffraction data, and solving crystal structures.

This facility supports simultaneous multiparametric analysis of many thousands of cells per second, enabling rapid analysis of complex cell populations using benchtop analysis flow cytometers. High-speed cell sorting services provide fast, objective and quantitative recording of fluorescent signals from individual cells combined with physical separation of cells of particular interest.

Whitehead Institute Flow Cytometry Core Facility
Whitehead Institute, Room 429
Contact: Kathleen Daniels
This facility is centered on flow cytometry, a laser-based technology employed in cell counting, cell sorting, biomarker detection and protein engineering, by suspending cells in a stream of fluid and passing them by an electronic detection apparatus. The facility has multiple instruments capable of single cell sorting and analysis.

W.M. Keck Microscopy Facility
Whitehead Institute, Room 447
Contact: Cassandra Rogers
This facility provides state-of-the-art light microscopy instrumentation and technical expertise to assist investigators in applying sophisticated imaging techniques in their research. The facility houses laser scanning and spinning disc confocal microscopes, a light sheet microscope, a range of wide-field microscopes, and a Transmission Electron Microscope (TEM). Training is required for access to all light microscopes and the TEM. TEM training is offered by the Koch Institute’s Nanotechnology Materials Core Facility.

Zebrafish Core Facility
Building 76-044
Contact: Adam Amsterdam
This facility was established to provide expertise to researchers wishing to utilize zebrafish as a model organism for any kind of research, along with professional daily care of the animals and aquarium systems. The facility can meet a wide range of needs, from one-off experiments of limited scope, to long-term maintenance of dozens of transgenic and/or mutant lines.
