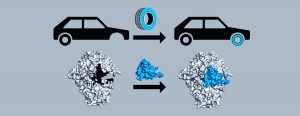
Bacteria use “spare part” proteins to repair damage and survive inhospitable conditions
Saima Sidik
Millennia ago, before the evolution of multicellular life, bacteria developed a group of proteins called glycyl radical enzymes to help them turn food into cellular energy. Glycyl radical enzymes functioned efficiently for millions of years until bacteria encountered a new hurdle: oxygen build-up in the atmosphere. Glycyl radicals are easily damaged by oxygen, and bacteria needed a way to continue to process nutrients under these new conditions.
In a recent study published in Journal of Biological Inorganic Chemistry, Sarah Bowman et al. from MIT provide structural evidence for how bacteria use “spare part” proteins to repair glycyl radical enzymes by replacing their damaged portions.
“Instead of degrading the whole protein and building a new one, bacteria make a small spare part protein that can bind to the glycyl radical enzyme and restore its function,” says Lindsey Backman, a graduate student in the MIT Department of Chemistry and co-author on the study.
Senior author Catherine Drennan, an MIT professor of biology and chemistry and a Howard Hughes Medical Institute Investigator, is leading the effort to characterize a glycyl radical enzyme called pyruvate formate lyase, or PFL, and its corresponding spare part protein, YfiD. Using a technique called nuclear magnetic resonance spectroscopy, her lab examined the shape of YfiD. Previous work revealed that upon oxygen exposure, part of PFL is cleaved off, leaving the enzyme unable to perform its chemistry. Then, using in silico modeling, the Drennan lab showed that YfiD fits neatly into the hole in oxygen-damaged PFL, replacing the missing piece.
“If you have a car and you get a flat tire, you don’t buy a new car, you just change the tire,” Backman says.
Although YfiD is one of only two spare part proteins that have been identified, Drennan suspects there are many more. It’s easy for researchers to mistake spare parts for the portions of proteins that they replace, and Drennan thinks that scientists have probably overlooked spare parts when analyzing the complex milieu of proteins found in cells.
Although identifying spare parts is technically challenging, Drennan thinks the applications for synthetic biology make it well worth the effort. Specifically, she’s interested in bacteria that live in the deep sea where there’s very little oxygen and use other glycyl radical enzymes to degrade hydrocarbons. Because they can break down hydrocarbons, these bacteria could be valuable tools for cleaning up oil spills. Oil spills occur on the ocean’s surface where there’s a lot of oxygen, so spare parts might be necessary to stabilize oil-degrading proteins under these conditions.
“The question always becomes, ‘what about the oxygen sensitivity?’” Drennan says, referring to these oil-degrading proteins. “What if we expressed a spare part to make them more stable?”
Backman began working on spare part proteins as an undergraduate participating in the MIT Summer Research Program, or MSRP, as did several of the study’s other authors, and Drennan says that this project’s success highlights the valuable role these students play in MIT labs. Now a full-time graduate student, Backman has teamed up with Drennan lab postdoc and co-author Mary Andorfer, and together they plan to continue characterizing the interaction between YfiD and PFL. They think their findings may be the first step in showing that spare parts are a common protein repair mechanism, and that characterizing them will add a new tool to the synthetic biology toolbox.
Citation:
“Solution structure and biochemical characterization of a spare part protein that restores activity to an oxygen-damaged glycyl radical enzyme”
JBIC Journal of Biological Inorganic Chemistry, online June 26, 2019, DOI: 10.1007/s00775-019-01681-2.
Sarah E. J. Bowman, Lindsey R. F. Backman, Rebekah E. Bjork, Mary C. Andorfer, Santiago Yori, Alessio Caruso, Collin M. Stultz, and Catherine L. Drennan.